Table of Contents
Even though I consider my place in science to be within the domain of microbiology, I am trained as a computer scientist. Similar to most others in life sciences that yield expertise in computation, our group started its journey in microbiology by mostly re-analyzing publicly available data. I have maintained both a ton of respect for those who had the vision to generate datasets of profound legacy and a healthy dose of envy. But all were OK. We did our best with what was available to us. Everything was beautiful and nothing hurt.
But as we learned more and more about microbial life through re-analyses of data others have generated, more and more questions started to appear in my mind for which no data were available to search for answers. Among all, there was this one question that was almost painful to think about since we didn’t have anything to go after it. We desperately needed a particular type of data that did not exist, and so our journey has started.
This blog post is about the question, the data, and the people who joined us towards generating them.
But why write a blog post on this rather than publishing your findings as a conventional article? Well. I really don’t have the ultimate answer to this question that is good enough to justify the time I am planning to put into writing this blog post. But I have some answers that motivate me. First, in a general sense, I am not happy with the fact that science is typically communicated by publications that do not really say anything about how it evolved over time. If a science project is a person, a conventional publication tells us only as much as a driver’s license would tell us about its owner. From the initial idea to the final data, a journey in science is a learning experience as a whole. Not only learning about science itself, but also its people. So this is my attempt to document the evolution of this one project I care a lot about.
Question
This section intends to only briefly explain the main theme of questioning that propelled this project yet it will still likely put you to sleep. But there is no gain without pain.
One of the most fundamental goals of our group has been to understand how do microbes respond to environmental change. We have studied this question in different habitats, and various temporal and geographical scales. Many of our investigations benefited from the TON of public data generated from environmental and host-associated microbial habitats. But in this particular case I was more interested in surface ocean microbes .. and change that takes place rather rapidly .. to which microbes respond beyond their genetic code. The ton of public data did not include anything to study that.
A relatively better introduction to this demands a better attempt to explain where I am coming from. Which is almost impossible to do well within the space constraints of an appropriately sized blog post, but I will offer a #YOLO version of it anyway.
All habitats on Earth force their residents to deal with change. Indeed at varying magnitudes, breadth, and speed, but constantly. Some ways for life to meet the challenge of change is to rely on the good ol’ boring means of evolution: a never-ending act of trial-and-error that influences the heritable genetic code in small ways to move an organism from one point in the fitness landscape to another. We have many ways to understand the impact and outcomes of such change through phylogenomics, pangenomics, population genetics, etc, strategies that allow us to dig into stories etched into DNA, such as high-throughput sequencing of genomes and metagenomes. But there are some kinds of change that are much faster than what conventional evolution and its excruciatingly unexciting, slow pace can keep up with. For that, there are other strategies by which life, especially microbial life, can take big strides in much smaller scales of time through mobile genetic elements, diversity generating retro-elements, hypervariable genomic islands, and so on. As the implications of such strategies to improve fitness are all present in the pool of -often- heritable genetic code associated with a microbial population or clade, they too are accessible to us to study through sequencing of DNA molecules. At even shorter timescales, living cells can simply adjust their ‘activities’ to respond to change without having to modify their gene pool. The first of such responses that come to mind is of course transcriptional regulation that can bump a population from one metabolic state to another by emphasizing different skills in its genetic repertoire. And of course this is not all. There are many more exciting and even unexpected responses take place when organisms are cornered to survive a sudden challenge. In the pool of such mysterious responses, some have been becoming more accessible to us to study over the recent years.
Cells have the means to respond to change even beyond their de facto capabilities defined by their gene content. A kind of response that is often invisible to us as they are inaccessible to popular sequencing strategies. A subset of such responses, and in my very biased opinion the coolest of all, involve transfer RNAs: the modest bringers of goods to the headquarters of translation, everywhere. Various properties of tRNA transcripts, such as their abundances and chemical modifications, can have a significant impact on protein synthesis in the cell, regulating what is translated, and how it is translated exactly. Therefore changes in tRNA transcript properties can shape the proteome and thus influence the overall fitness of cells that they are serving.
If you are a microbial ecologist, it is more likely than not you are grossly underestimating the versatility of tRNAs. If you are in the mood of losing a bet almost instantaneously by disagreeing with me, you may proceed to quickly skim through the subsections of this lovely paper by Medha Raina and Michael Ibba. I contextualize this with this analogy: if the genome is a cookbook and the ribosome is the chef that reads that book and prepares dishes of all sorts, there exists a class of epi processes that influence transfer RNAs, through which the cells can prioritize the translation of certain proteins independent of levels of transcription, or even let the chef deviate from the recipe through mistranslation events and produce versions of proteins the cookbook does not described. This profoundly under-explored and a very exciting layer of biology has been steadily gaining traction thanks to advances in technology. If you are interested to dive into that literature from a random part of it, here is a fascinating paper on this topic, a very recent study that can give you even more insights, and an excellent short review that contains many relevant citations.
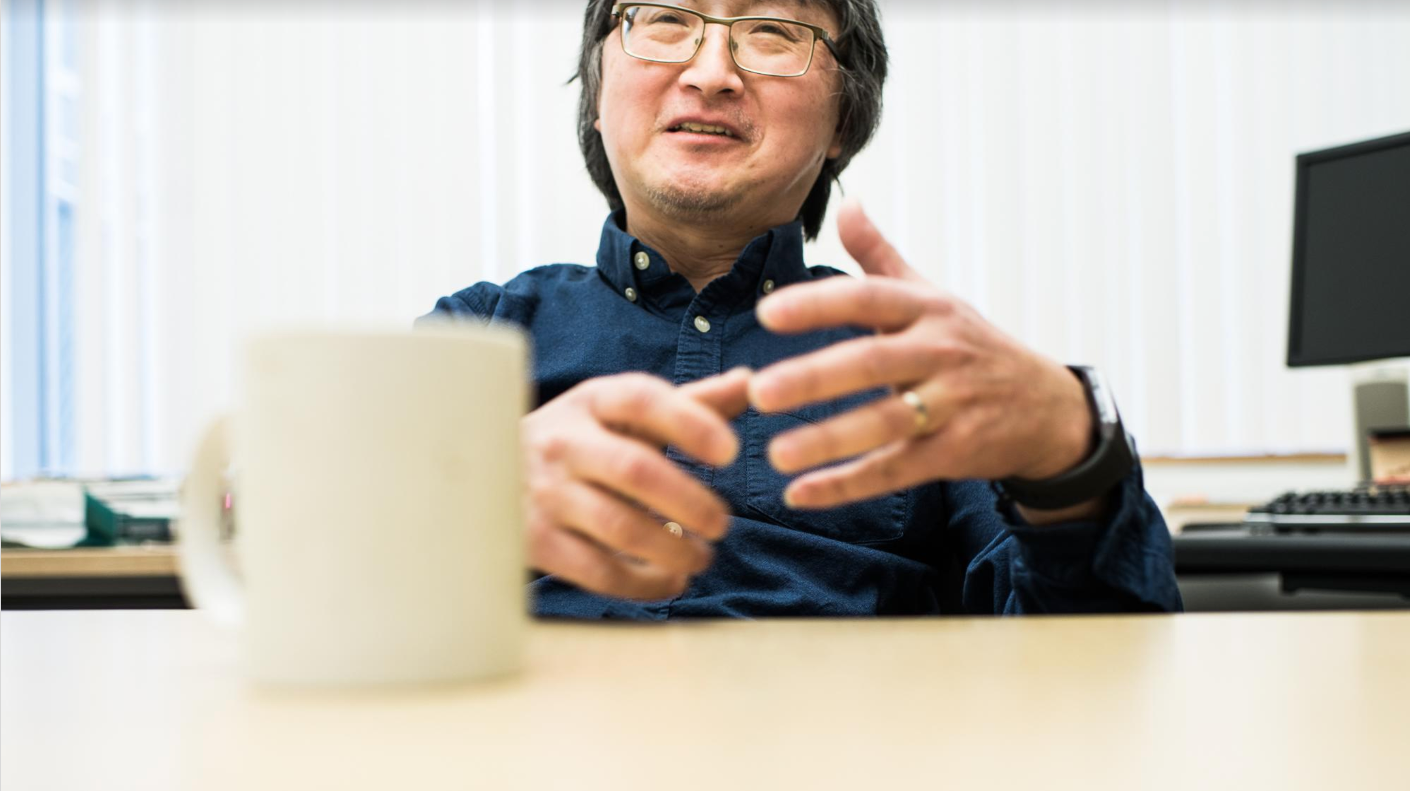
I was introduced to this entire field a few years ago by Tao Pan.
Tao is a professor of biochemistry at the University of Chicago whose expertise lies in RNA biology. We met in 2016 during a graduate student event. He sat next to me during the dinner, and told me that his group developed a new molecular strategy to sequence tRNA transcripts in an high-throughput fashion (which historically has been an extremely difficult task due to their rigid secondary structures), but the FASTQ files they were getting from Illumina machines contained way too much information for them to deal with. Having worked with ribosomal RNAs for a long time, I thought dealing with the data generated from tRNAs would be a piece of cake for me simply because,
query ...: tRNA
|||
target ..: rRNA
So I replied, “say no more fam”.
Years later I was going to find myself remembering that moment and contemplating how my life is littered with moments of ignorance like this that somehow both embarrassed me and helped me in the long run.
Well. Dealing with tRNA transcripts of course proved to be EXTREMELY difficult. Much much more difficult than dealing with rRNA amplicons, in fact. Now when I think about tRNAs, I feel like rRNAs have always been too good to be true purely from a data analysis perspective. rRNAs were dogs: molecules that give us everything they can within the boundaries of their limitations. They were so good, we didn’t deserve them. In contrast, tRNAs were hyenas: they may look and sound like rRNAs to untrained ears and eyes, but they would come in groups, tear you up with thier volatile personality, and proceed to start consuming you while you protest. I was not trained enough to know this in 2016. I am now. And if you are a computer scientist or a statistician who is interested in developing novel algorithms for some very difficult and novel problems you will likely never be able to solve to your satisfaction, let us know. We have them.
But things were not this grim at the beginning. In fact, our initial interactions with Tao resulted in the following study in 2018:
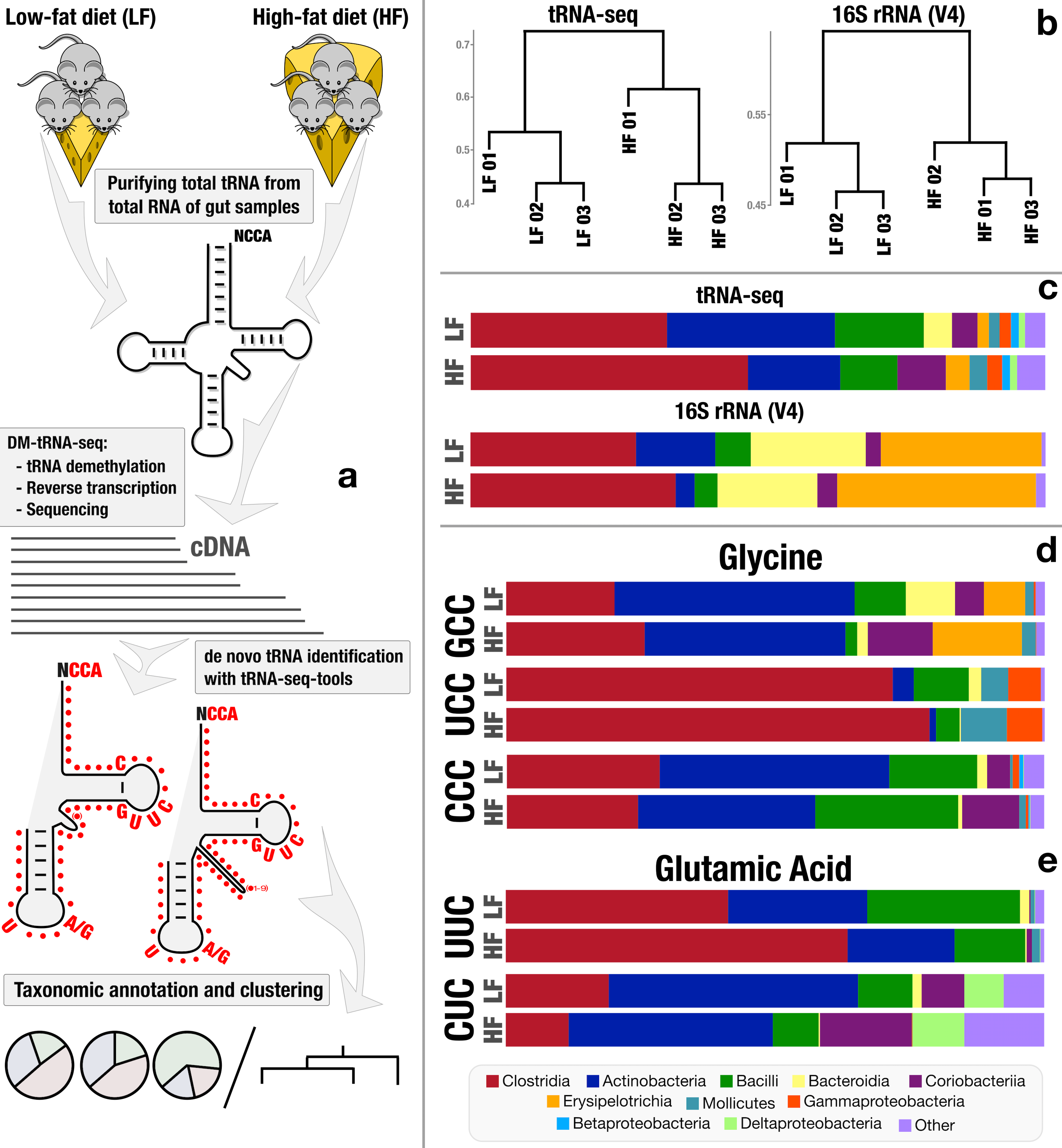
- Reveals taxon- and diet-dependent variations in tRNA modifications, and provides first in situ insights into 'metaepitranscriptomics' through tRNA gene expression dynamics and post-transcriptional modifications.
- "New RNA sequencing strategy provides insight into microbiomes", by Matt Wood.
- "RNA sequencing offers novel insights into the microbiome", by Liji Thomas, MD, and Kate Anderton, B.Sc.
This pilot study was the very first application of the fancy tRNA sequencing strategy to a complex, naturally occurring microbial habitat in the mouse gut. In this study we observed that the sequencing of tRNA transcripts from complex microbiomes could yield critical insights into microbial responses to their environment which we could not quite learn from other popular and affordable data types. In addition to that, this study helped me appreciate the power and the intricacies of working with tRNA transcripts. I was sold. But there was A TON of molecular and computational work to do if we really wanted to use this ‘metaepitranscriptomics’ approach effectively, and most importantly, make it accessible to others.
Our desire to advance this work was too high-risk for federal funding. But luckily, under Tao’s leadership, we received a substantial support from the W. M. Keck Foundation to push this research forward. During the last three years Tao Pan and members of his group, especially Chris Watkins and Chris Katanski, has been working tirelessly to improve molecular biology that underlie tRNA sequencing to achieve higher throughput and higher accuracy with less input biomass so we could apply this strategy to environments outside of the gut. In parallel, my group was supposed to work on the computational challenges and the integration of tRNA sequencing with other ‘omics strategies. I was extremely lucky to start working with Sam Miller to do that. Sam has been addressing unspeakable computational challenges one by one to make sense of tRNA sequencing results. If you can read Python code, just take a look at the intricacies sorted out in this module that is supposed to simply identify and characterize general features of tRNA molecules from tRNA sequencing data. Sam’s thousands and thousands of lines of code to tame the sequencing data from tRNA transcripts have already resulted in multiple programs such as anvi-trnaseq and anvi-convert-trnaseq-database that are just waiting to appear in a publication that demonstrates their utility and place in microbial ecology.
Sooo, the long story short, I was realizing in 2019 that (1) the sequencing of tRNA transcripts had the potential to take environmental microbiology to new pastures, (2) our molecular approaches that simplify library preparation stages were improving for applications of tRNA-seq in complex and low-biomass environments thanks to Tao, Chris K, and Chris W, and (3) our computational strategies to study tRNA abundance and chemical modifications through resulting data were maturing in Sam’s talented hands. Needless to say, I was dying to see whether the application of these advances to marine systems could bring us closer to investigate a slightly more refined form of the general question I mentioned at the beginning of this chapter to go after how do surface ocean microbes respond to environmental change BEYOND the heritable genetic code in very short scales of time?
It was clear that we had to do some high-throughput tRNA sequencing on some surface ocean samples. But we needed two more things to come together to generate the data we needed: financial support, and motivated people.
Finances
Finding financial support for high-risk ideas is extremely difficult. Especially for early career scientists who do interdisciplinary work. For instance, I quickly learned that funding decisions are inevitably driven in part by the reputation of the investigators who apply for funding. The contribution of reputation to these decisions is not something that is very obvious to those of us who never sat through an entire NSF grant review panel or those of us who sat through too many of them… But the grant applications of all early career scientists, especially those who were not trained in superstar labs or coming from superstar institutions, do suffer from not having a reputation in the field. And who can really blame the reviewers who are given the responsibility by the funding agencies to help determine how to partition our scarce financial resources if they are biased towards groups that have demonstrated successes? So that’s that. Being in between multiple disciplines does not help either. I wish I had the courage to share some of the comments I have received for our papers or grants we sent to NSF and NIH over the years. In addition to not having a reputation, I often was neither enough of a computer scientist for computer scientists, nor I was enough of a microbiologist for microbiologists. My name was Neitherherenorthere, PhD, which translated to being too risky to support through federal funds.
So from where I could find support for this ambitious and high-risk project? My mentors at the Department of Medicine at the University of Chicago, and especially my section chief David Rubin, have always been there with an inexhaustible confidence in me and support for my random ideas. But it would have been too embarrassing to ask for the section of ‘Gastroenterology, Hepatology, and Nutrition’ to support marine research. If I was going to do this, I had to do it right.
While I was looking around to find an appropriate opportunity, I learned that the Simons Foundation was quite welcoming to new investigators. So in 2020 I applied for the Early Career Investigator in Marine Microbial Ecology and Evolution Awards. This was an excellent award mechanism that aimed to help investigators who wished to advance our understanding of marine microbial ecology and evolution, and interdisciplinary ideas were welcome.
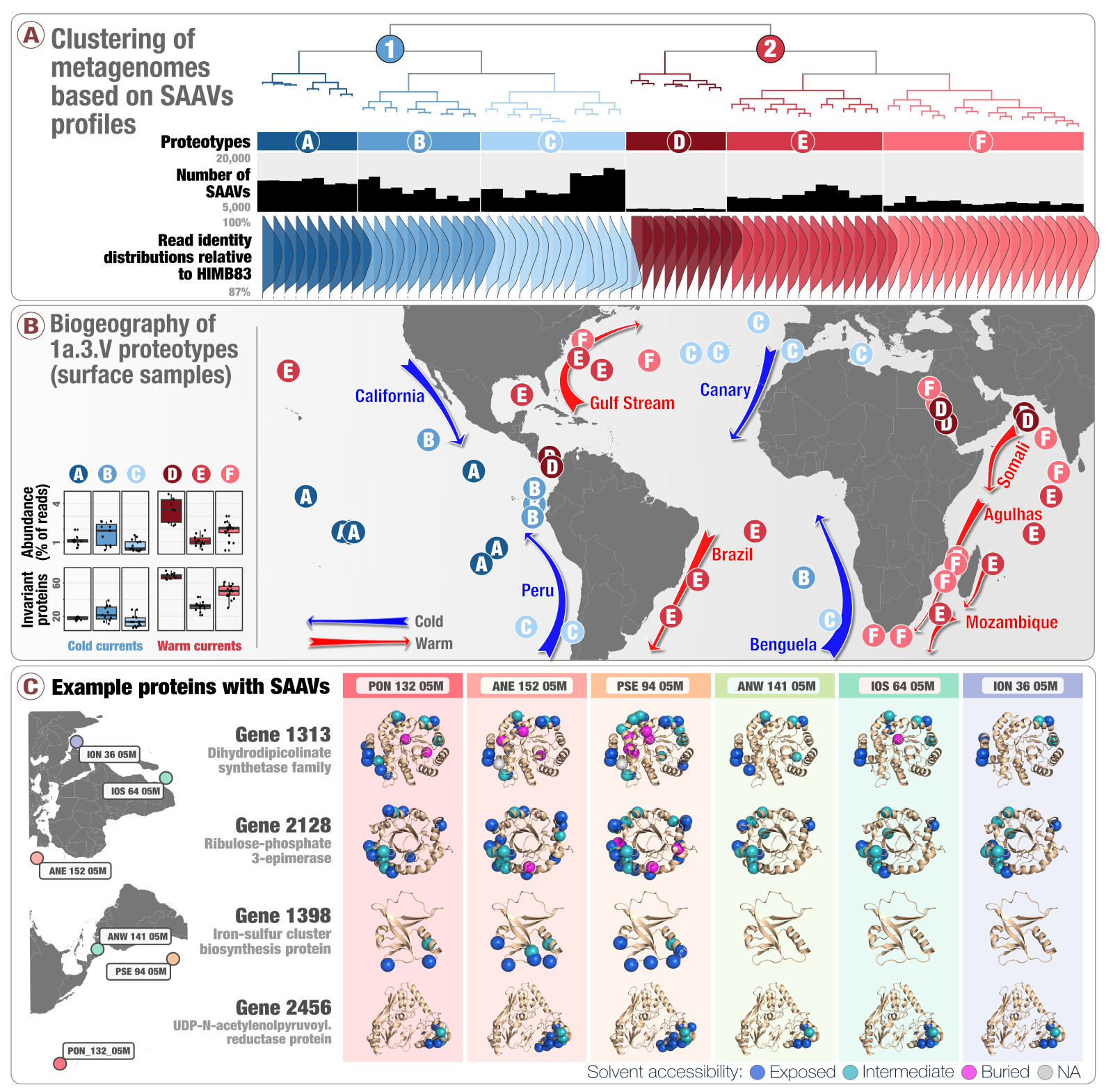
The proposal was building on our previous work on microbial evolution in marine systems that was co-led by Tom Delmont, a creative microbial ecologist who pushed me into the world of ‘omics, and Evan Kiefl, a talented researcher who has become one of the most prolific contributors of anvi’o during his graduate work:
- A first attempt to link population genetics and the predicted protein structures to explore in silico the intersection beetween protein biochemistry and evolutionary processes acting on an environmental microbe.
- An application of metapangenomics to define subclades of SAR11 based on gene content and ecology.
- Reproducible bioinformatics workflow is here. Reviewer criticism and our responses are also available.
My proposal suggested that the next step was to bring in tRNA sequencing into the mix. It was ambitious, but also transparent about what we didn’t know. I literally put the questions I had in mind in the proposal as questions: What can we learn from the high-throughput sequencing of tRNA transcripts from surface ocean microbial populations? How can we use metaepitranscriptomics in conjunction with metagenomes and metatranscriptomes to create a more refined understanding of the ecology and environmental responses of major marine microbial clades? I also argued that the successful completion of this proposal could offer high-resolution insights into microbial strategies for survival and adaptation by extending the current ‘omics arsenal available to environmental microbiology.
I remember exactly where I was when I got the official notice from Marian Carlson, the director of Life Sciences at Simons Foundation, that my proposal was selected for funding. It was on March 13, 2020. I responded to the news and mentioned that my group and I were very excited for the next three years. Ironically, this was exactly two days after the World Health Organization had declared coronavirus a pandemic. This was only weeks after I was named as one of the Research Fellows of the Sloan Foundation in Ocean Sciences. So life had other plans for all of us, but thanks to support from the Simons Foundation and Sloan Foundation, I now had the financial support and thus the responsibility to study the surface ocean metagenome, metatranscriptome, and metaepitranscriptome.
The next challenge was to actually sample the ocean. Which required bringing together a group of people who would be willing to take part in this journey.
People
As fundamental expertise on the Chicago side for a future sampling saga, I had by my side my colleagues Karen Lolans, an expert molecular microbiologist in our group who has been mastering high-molecular weight DNA extraction protocols and preparation of tRNA sequencing libraries, and Jessika Fuessel, a marine biogeochemist with extensive expertise in sample collection and processing from oceans.
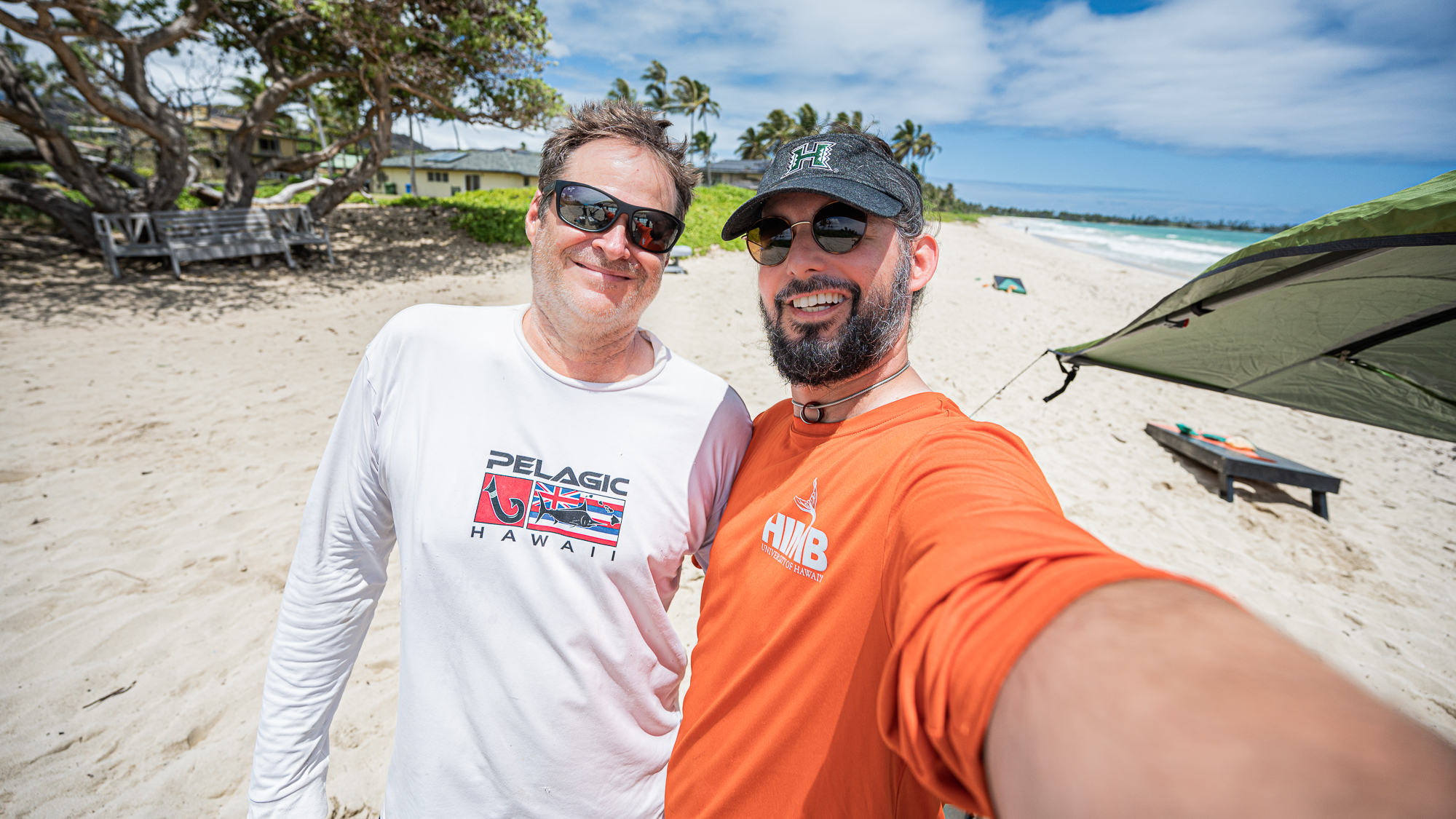
But we could not do this without substantial help. One of our long term science partners, Michael Rappé and his group at the Hawai’i Institute of Marine Biology (HIMB), were aware of this proposal since its conception. They graciously agreed to work with us on this.
The initial plan Mike and I concocted and run through Tao was simple:
(1) Go to Hawai’i, where the Rappé group had access to both coastal and open ocean stations, (2) collect samples every hour for 48 hours from two stations and immediately process them for short-read metagenomics, long-read metagenomics, metatranscriptomics, and tRNA-seq, (3) generate sequencing libraries back in Chicago, (4) analyze data.
I call this the ‘PI Plan’. A more of a blue-sky thinking, or like an act of wishful thinking, and not to be confused with the ‘Real Plan’. Which requires addressing a countless number of theoretical and practical considerations for that wishful thinking to turn into actual high-quality data generated in a safe environment. In our case the 48 hours continuous sampling meant at least two teams to work around the clock for two days to drive boats, collect, process, and store samples. So the real plan required much more than the PI plan. Figuring out and testing in advance the sampling strategy, the sampling equipment, and protocols for filtering seawater and extracting DNA and RNA molecules were all parts of the real plan. The real plan required putting together essential chemical and physical consumables, ordering them in a timely fashion to ensure that they could make it to the sampling effort in time. The real plan required finding volunteers to work in shifts, organizing people on a time table, making sure they had the means to make it to their shift, and coming up with contingency plans for worst case scenarios. The real plan required acquiring traveling and accommodation permissions from institutions, keeping an eye on the weather to determine the best timing of the start of the sampling, and designating certified boat drivers for the prime time. The real plan was not done yet. It also demanded finding people places to sleep in between shifts, figuring out how to feed and hydrate people during their shifts, and preparing sample names, and sheets, and labels, and finding cars, pens, filters, filter caps, nylon nets, fuel for cars, fuel for boats, coolers, fans, carboys, hats, headlamps, sunscreens, life vests, gloves, trays, ice packs, freezer storage spaces, and more. The real plan was real, and it could be put in motion only by those who were truly passionate about science to a degree of geniune selflessness.
Many people contributed to the emergence of the real plan. But ultimately the initial idea was going to grow its wings thanks to Kelle Freel (a post-doctoral researcher who uses cultivation approaches and sequencing of environmental samples to study marine microbial communities in the surface ocean as well as the in the basaltic crust below the seafloor), Sarah Tucker (a PhD candidate interested in marine microbial ecology and evolution and combining field and laboratory techniques to further understand microbial metabolisms and their important roles in biogeochemical cycling), and Jessika Fuessel, who worked very closely with them.
The rest of us, Mike Rappé, Karen Lolans, Evan Kiefl, and myself helped with the initial planning, too. And in fact even more people joined us during the actual sampling. But the true weight of turning a vague idea into reality did sit on the shoulders of Kelle, Sarah, and Jessika. Which made me think. A lot. About the fact that from all disciplines of science papers with data emerge thanks to the hard work of so many people who often are, at least in the long run, known as et al.
I admit here in embarrassment that while observing the work ethic and dedication of those who joined this effort, I realized that I never fully appreciated the true cost of generating data for those who will most likely not going to get the credit they deserve in full. I have strong opinions on public data: I believe data that are paid for by public resources or data that are used to make public statements must be made public with no strings attached. Which is not as smooth as it should be despite progress. But another aspect of public data that I have not paid attention to as much as I should have is the need to properly acknowledge the invaluable contributions and hard work of those who helped materialize science by turning mere ideas into plans and data when the linear order of names in publications that build on their efforts have only two prime positions to highlight.
Our group respects data generators and we acknowledge them and their influence on our work with statements like this one:
(...) we are indebted to the scientists who made this study possible by generating the genomes and metagenomes, and making them publicly available.
We are indeed indebted to them. But if I have to be brutally honest for a moment, ‘respecting’ or ‘acknowledging’ data generators, whose names in many cases are collapsed into et al statements, is similar to walking out of a supermarket with a paper bag and feeling content for being on the right side of history as far as the Great Pacific garbage patch is concerned. Unless these acknowledgements actually resolve to action that impacts their career. Here I probably finally converge with many scientists who already have realized the critical importance of recognizing those who contribute to science beyond first and last authors. I am sorry for being so late. I will do my best to catch up.
Putting that discussion in the back burner and coming back to our smaller reality, after much work, we had a real plan: Evan, Jessika, and I were going to be in Hawai’i on August 16, 2021, and the weather permitting, we were going to start the 48-hour sampling marathon on August 18, 2021.
Sampling
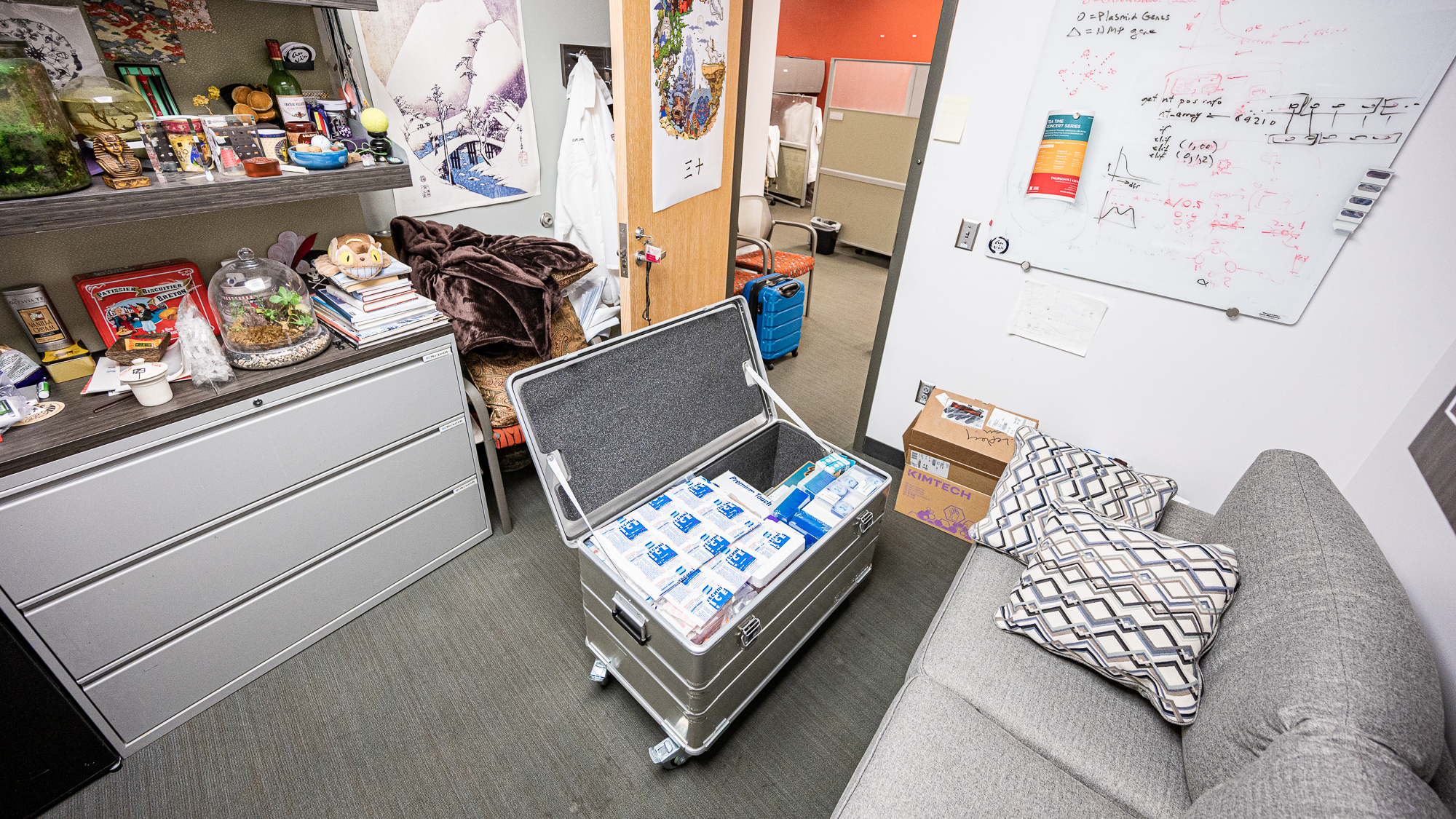
Matching the increasing number of emails that were flying across the Pacific Ocean as Sarah, Kelle, Jessika, and Mike were trying to put together a list of consumables and equipment we were going to need during the sampling, there was a Zarges box in my office at the University of Chicago that was filling up with sterivex filters (to trap unexpecting microbial cells from seawater), Cryo-Babies (labels that are designed to withstand boiling, freezing, and liquid nitrogen storage without peeling to put on tubes), periplastic pump tubings (in case something goes wrong with tubings we can continue filtering without interruption), RNALater solutions (salts that protect RNA molecules from degrading (which, we recently learned through an experiment, protects also the DNA molecules from fragmentation (wink wink high-molecular weight DNA extraction enthusiasts))), and gloves, and labels, and caps, an nylon nets – all sorts toys for wet-lab people.
August 16
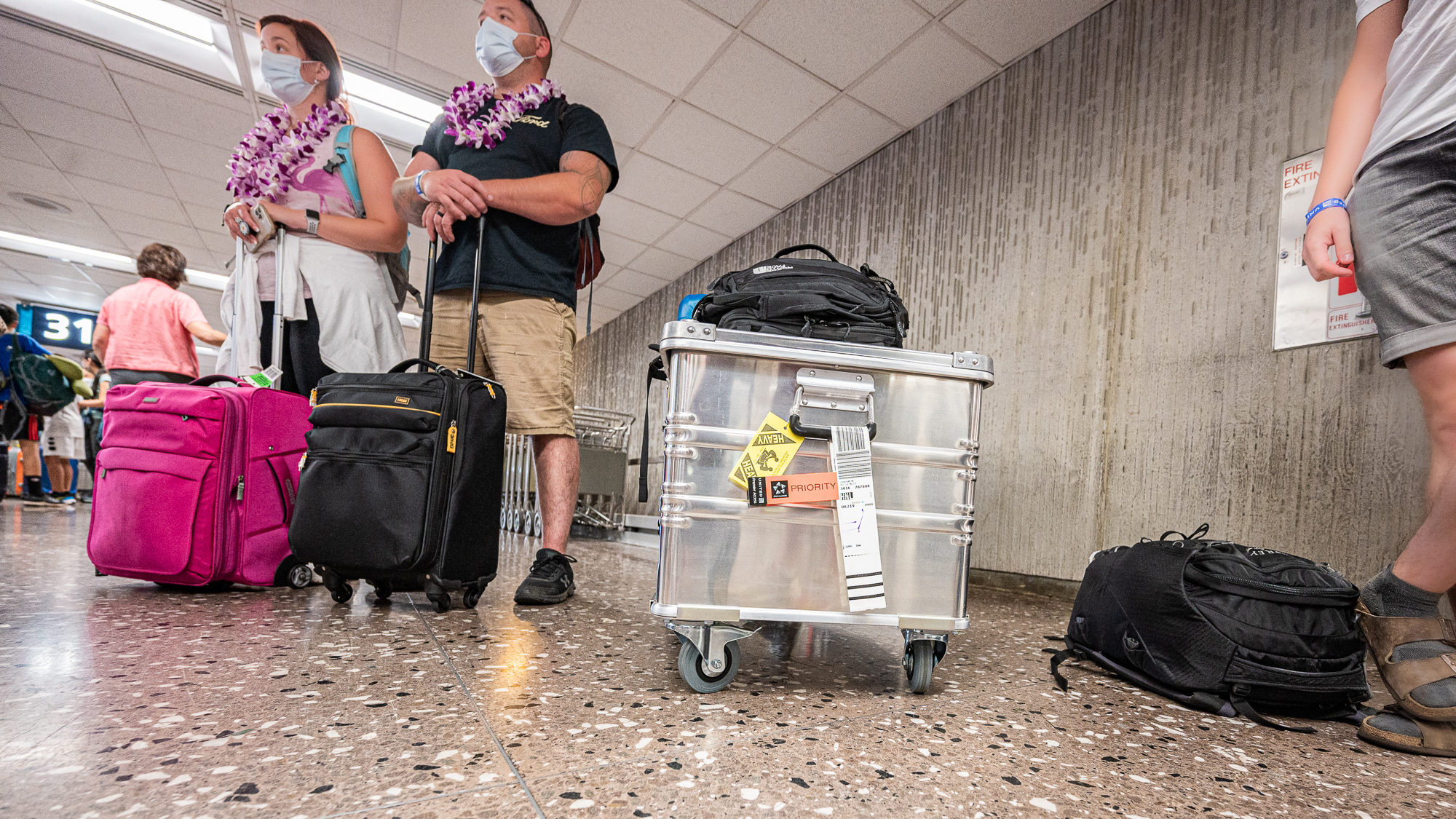
In the morning of our flight to Hawaii, the Zarges box was proudly displaying its contents with absolutely zero appreciation of the amount of energy it was going to take to bring it to its final destination that was more than 4,000 miles away.
Our flight was around 10am, but after all the preparations for this sampling, being late to the airport was too much of a risk. Despite the protests of the remaining members of this fellowship, we were in our cab headed towards the airport at precisely 5:00am. Our cab did travel at speeds we were familiar from Star Trek and made it to the airport in record time. We were there around 5:20am (successkid.jpg).
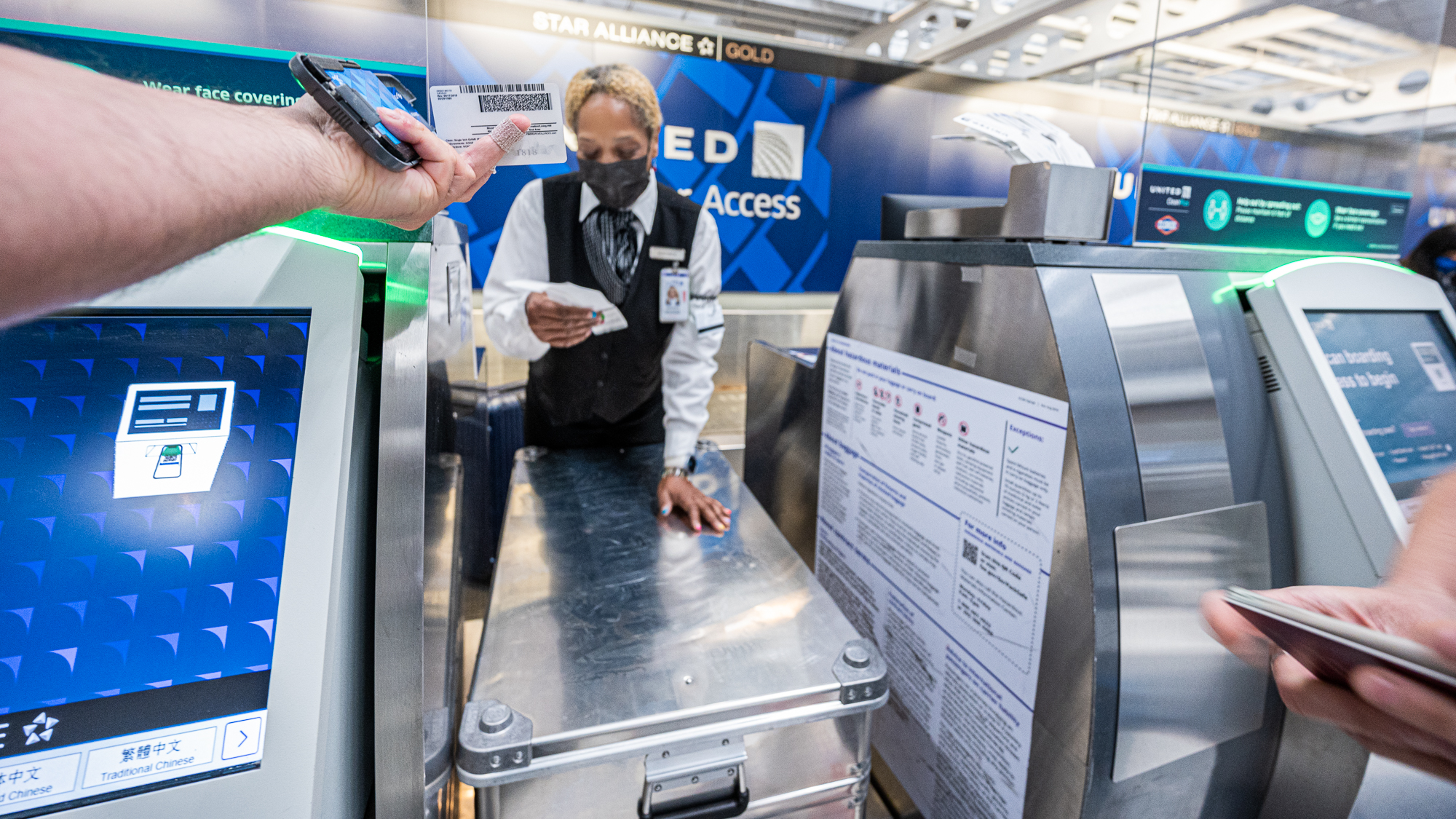
As someone who travels light, I have very little experience with checking in luggages for flights. And as someone who is extremely unlucky, I have a lot of experience with things going unexpectedly wrong with my interactions with the airport people. So my brain was already cooking up some dark scenarios. Perhaps they were going to say our Zarges box was too heavy. Or too ugly. Or too full. And we were not going to be able to bring it to Hawai’i. I already was having hypothetical conversations with the United representative preparing myself to that hypothetical moment where they were going to refuse to take our box to the airplane. My acid reflux was helping quite a lot.
We can’t take the box? OK. I see. But listen – I know you know all about the canonical role of transfer RNAs in protein synthesis, but did you know that the last few years witnessed so many groundbreaking studies that revealed additional roles of tRNAs in translation regulation? And then there are all these microbes in the ocean, right? It is all like really very important AND THE ANSWER IS IN THIS BOX BEFORE YE. Well, not in this box exactly, but if you let this box go into that plane, we can tell you MUCH MORE about this in our pre-print that will likely appear on bioRxiv in like 4 years from now. Yes, you MAY take my drivers license hostage until then. Yes, that’s me in the picture. Yes, my hair has a lot more whites now. Yes, I do look like as if I was high when they took that pic …
My preparations were all for nothing. The representative did not even flinch seeing our box. I was both happy and disappointed.
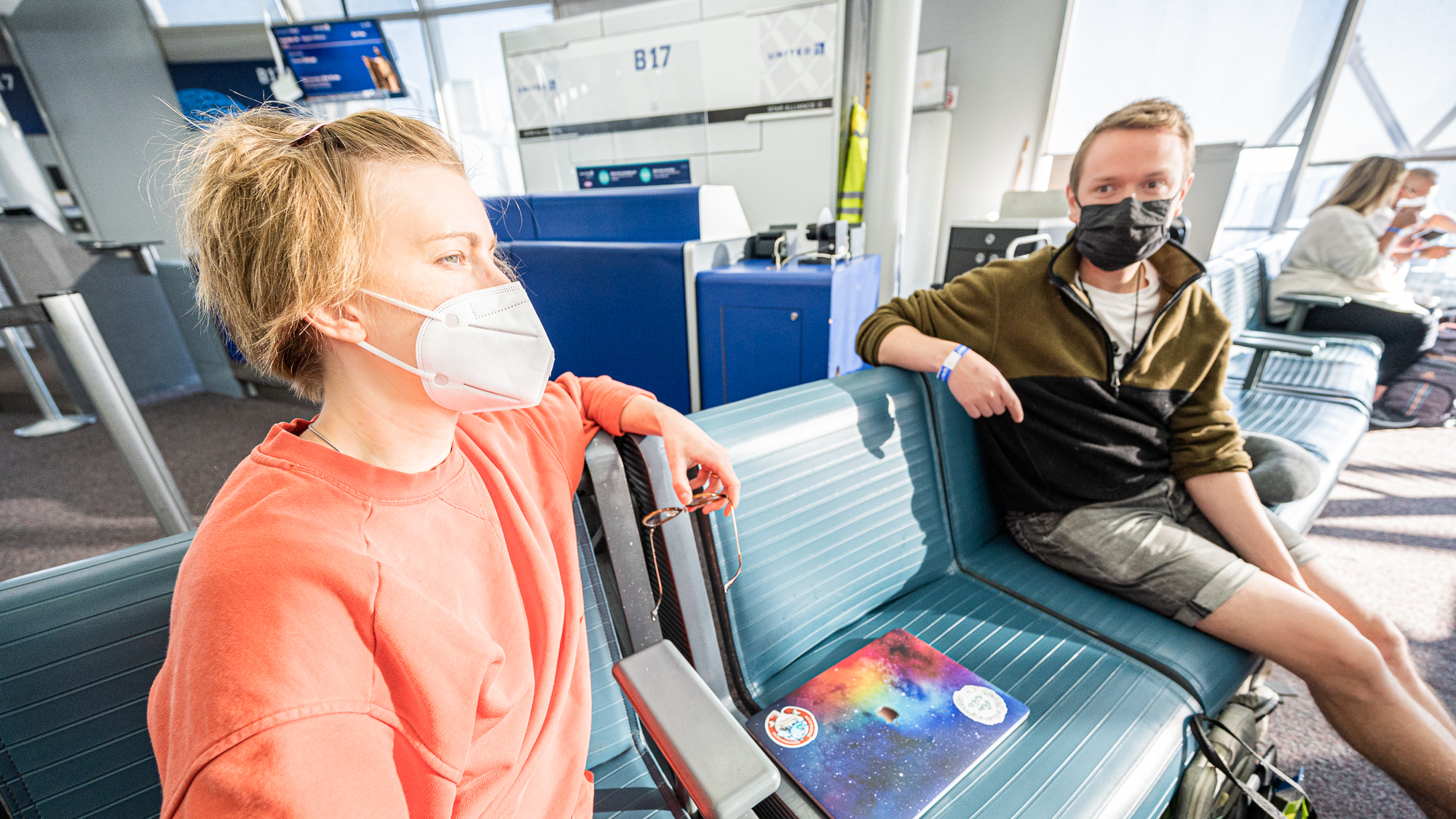
So the fellowship had made it to their gate .. a few hours before the boarding. After about an hour of sitting on our chairs, both Jessika and Evan were slowly realizing how worthy it was to wake up at 4am to be here: our journey to Hawai’i had started with so much joy that their masks were barely containing their smiles:
Althought it was filled with so much love, there was no traditional lei greeting at Honolulu for the Zarges box when we finally landed in O’ahu.
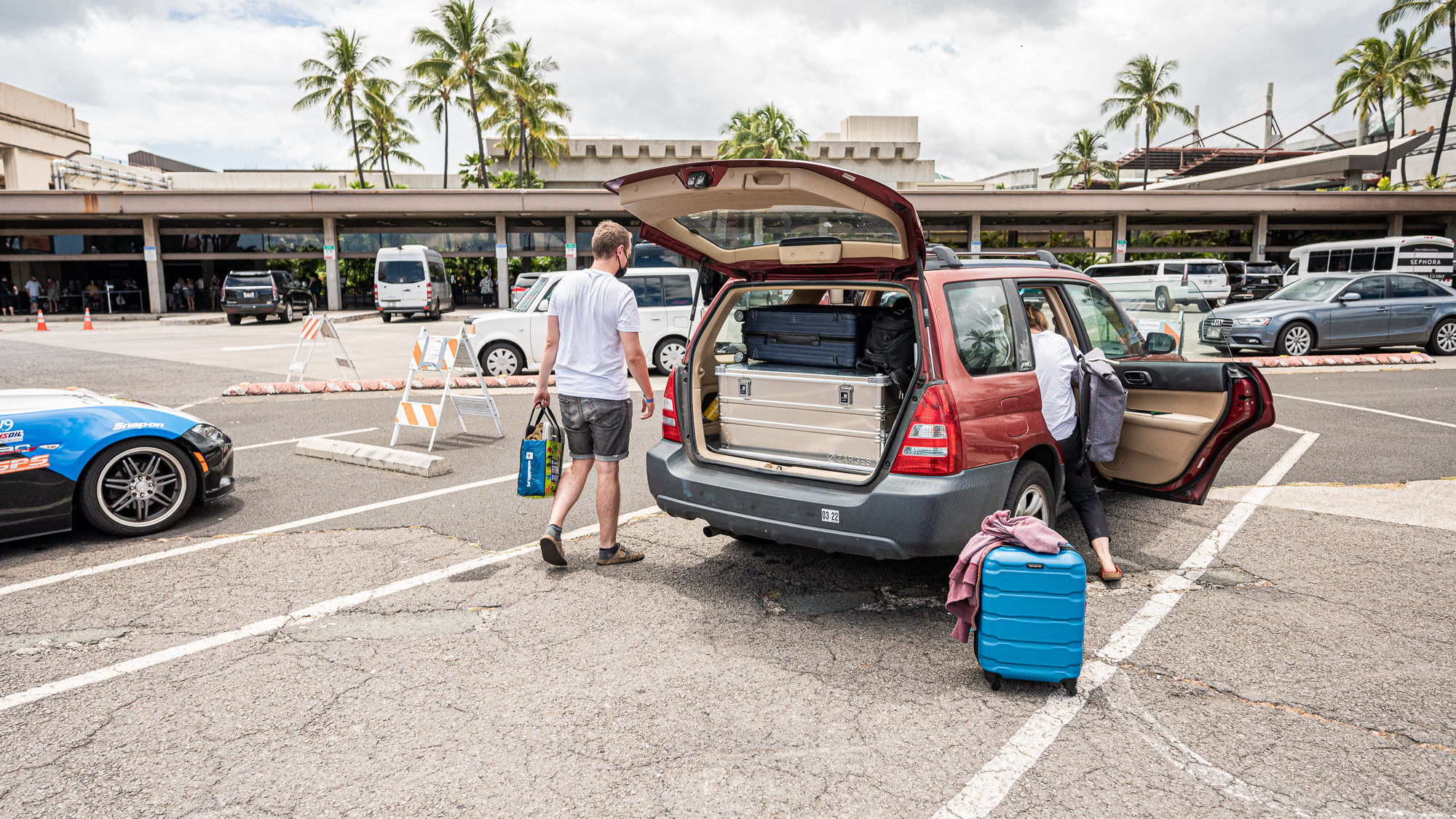
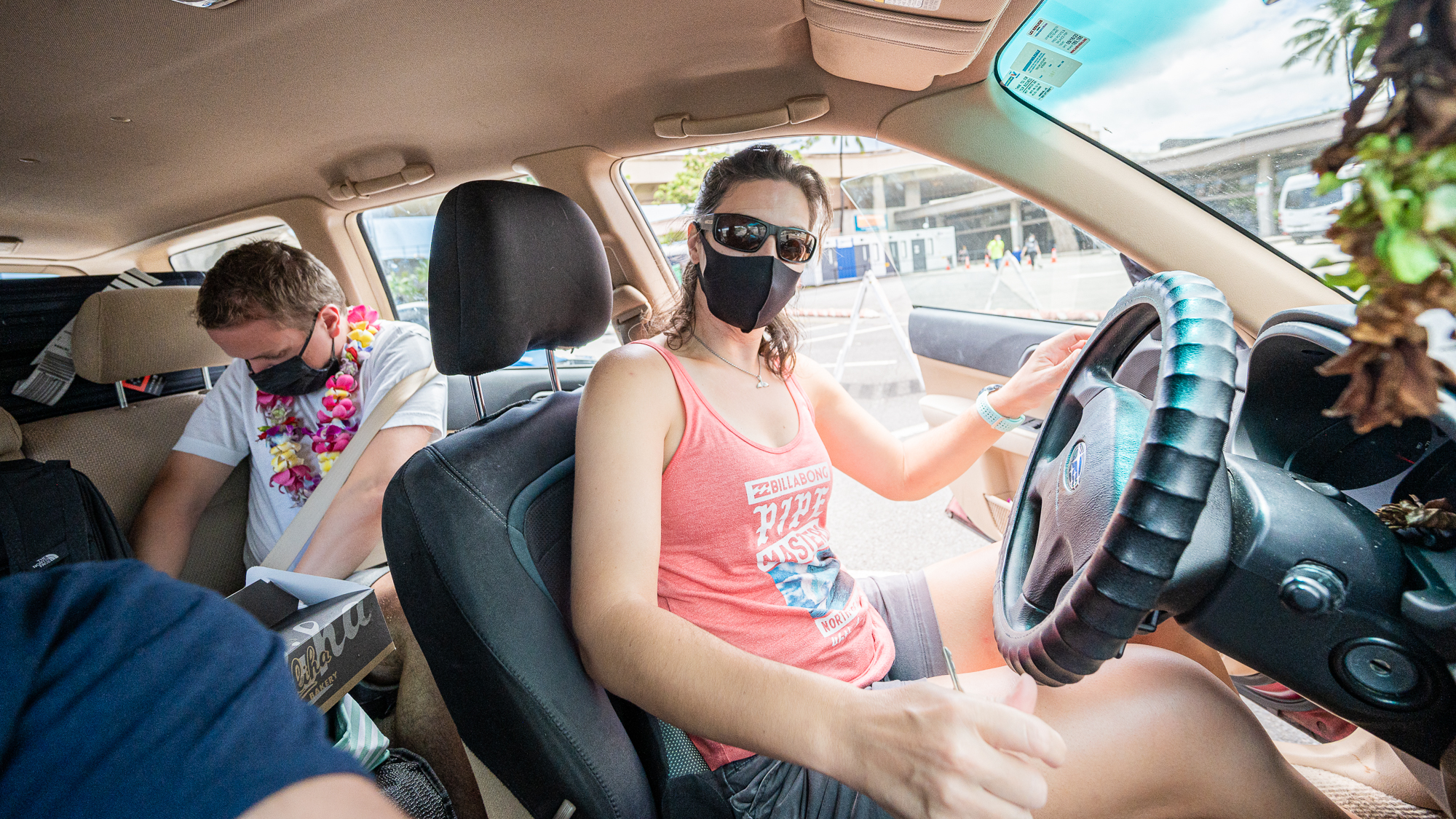
After a short wait, Kelle arrived to collect us from the airport, AS PLANNED.
She had brought not only lei for all of us, but also a box full of green tea cream puffs and poi doughnuts from Liliha Bakery, which was not a part of the plan. But it was the best welcome imaginable.
Our destination was Hawai’i Institute of Marine Biology, a world renowned institution focusing on understanding and conserving tropical marine ecosystems, and the professional home of the Rappé group.
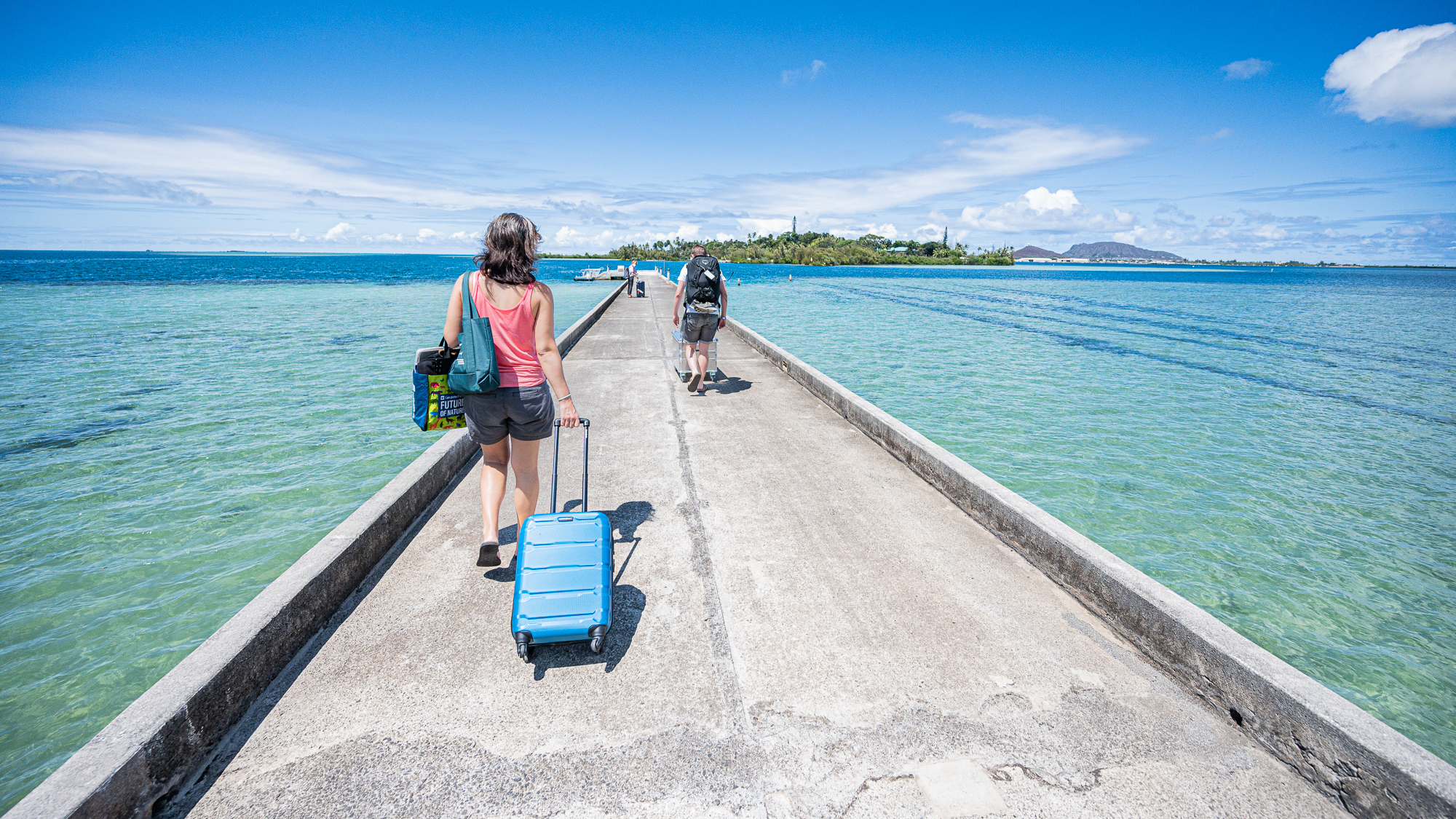
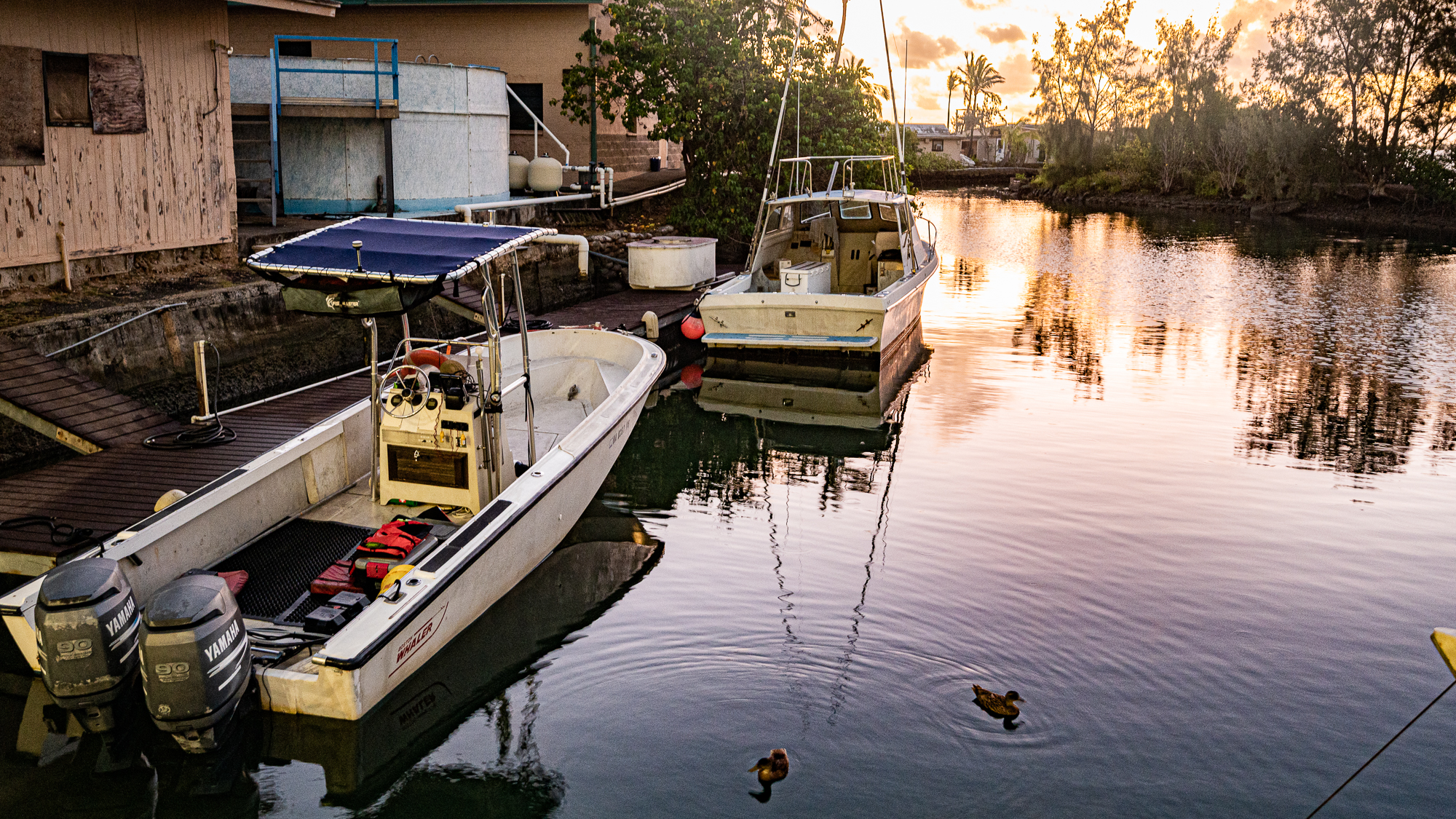
HIMB is on the Coconut Island (Moku o Lo’e). Which is a tiny island that is just minutes away from O’ahu. Even though it is right there, the only access to the island is through a boat like most islands. So it is a proper island. Just tiny. We pulled our belongings to the dock. But not Evan. Evan pushed the Zarges box by yelling at it and it was very effective:
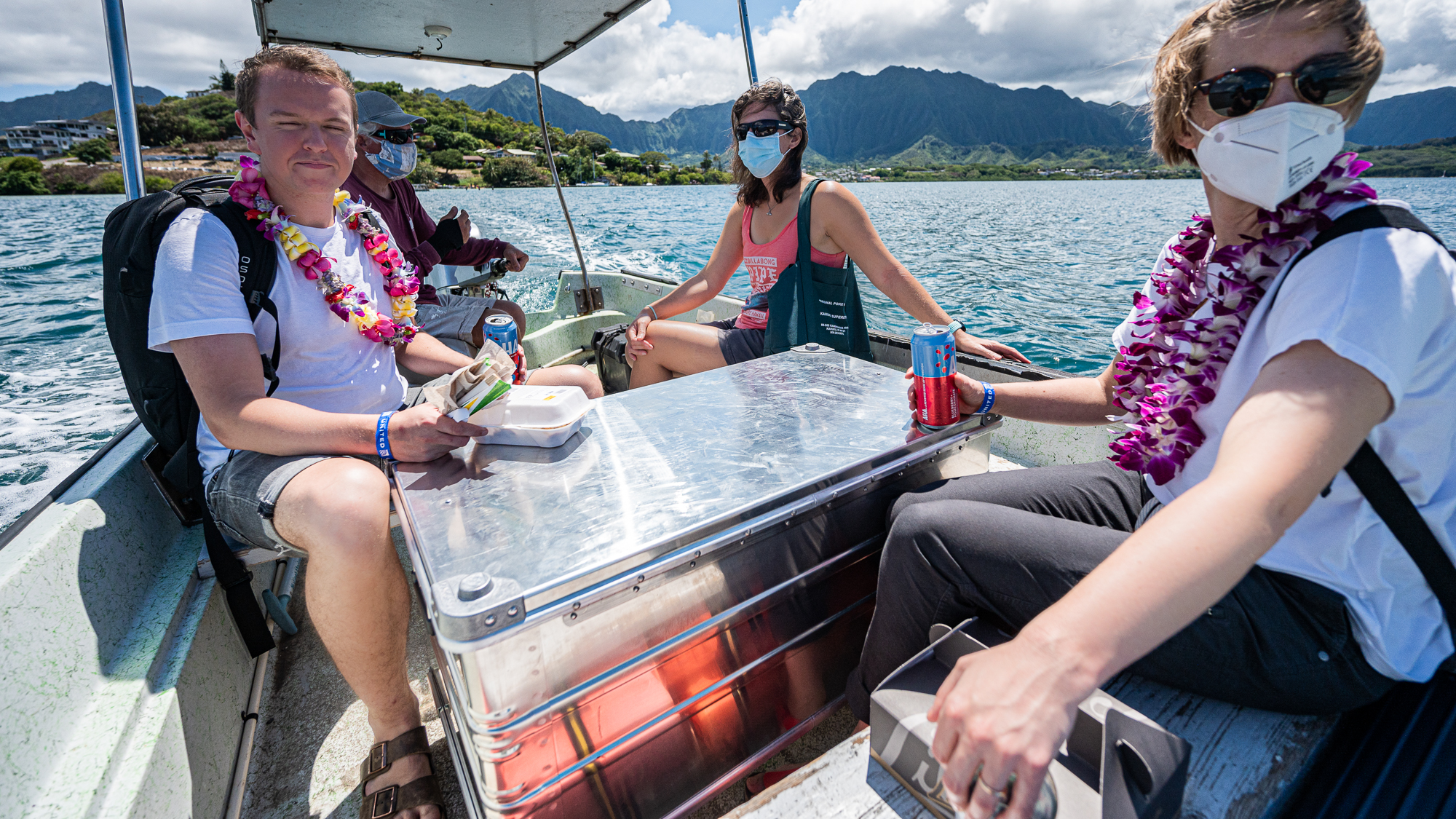
Moments later, we were on our short and sweet boat ride to settle in our housing.
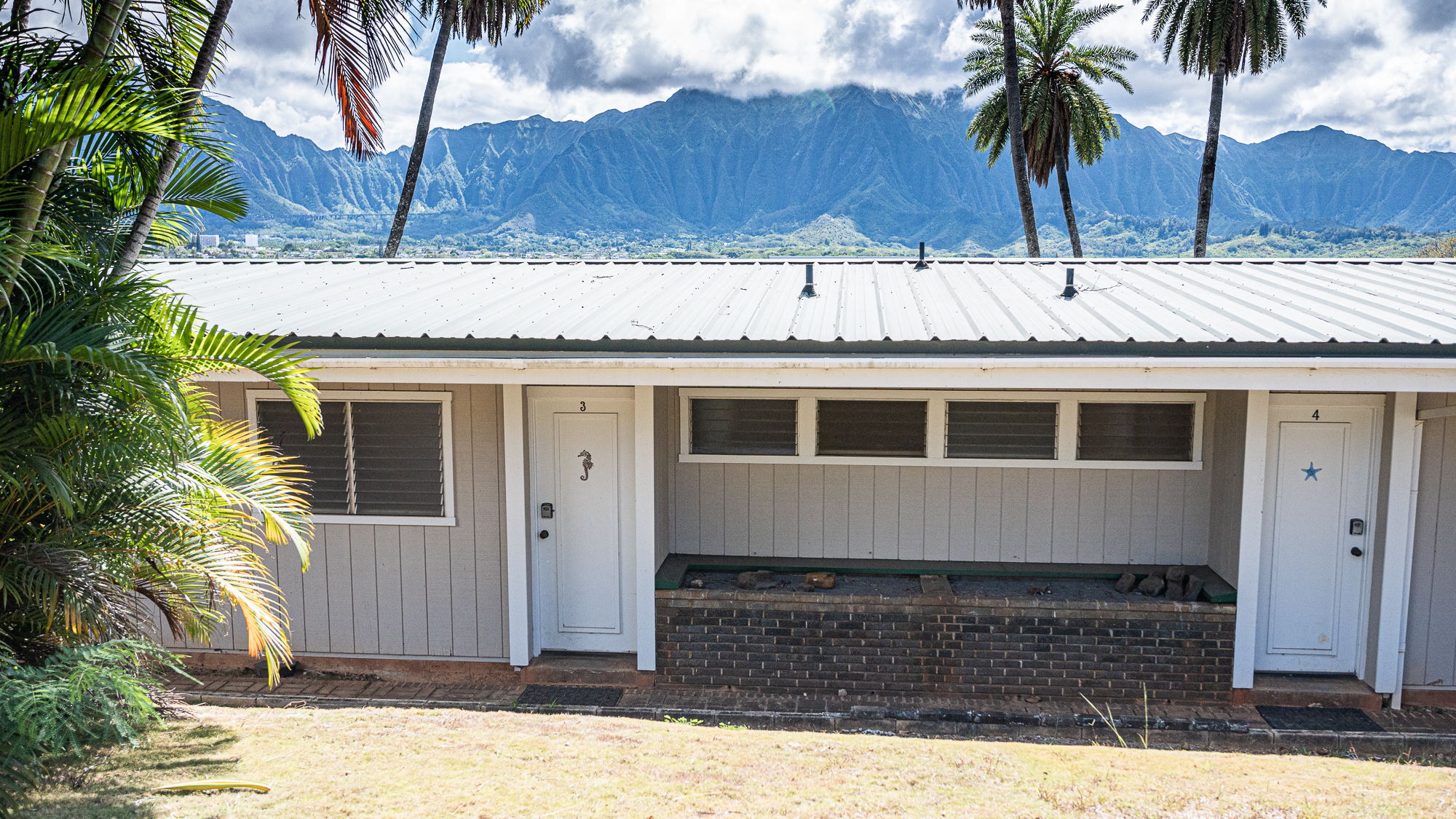
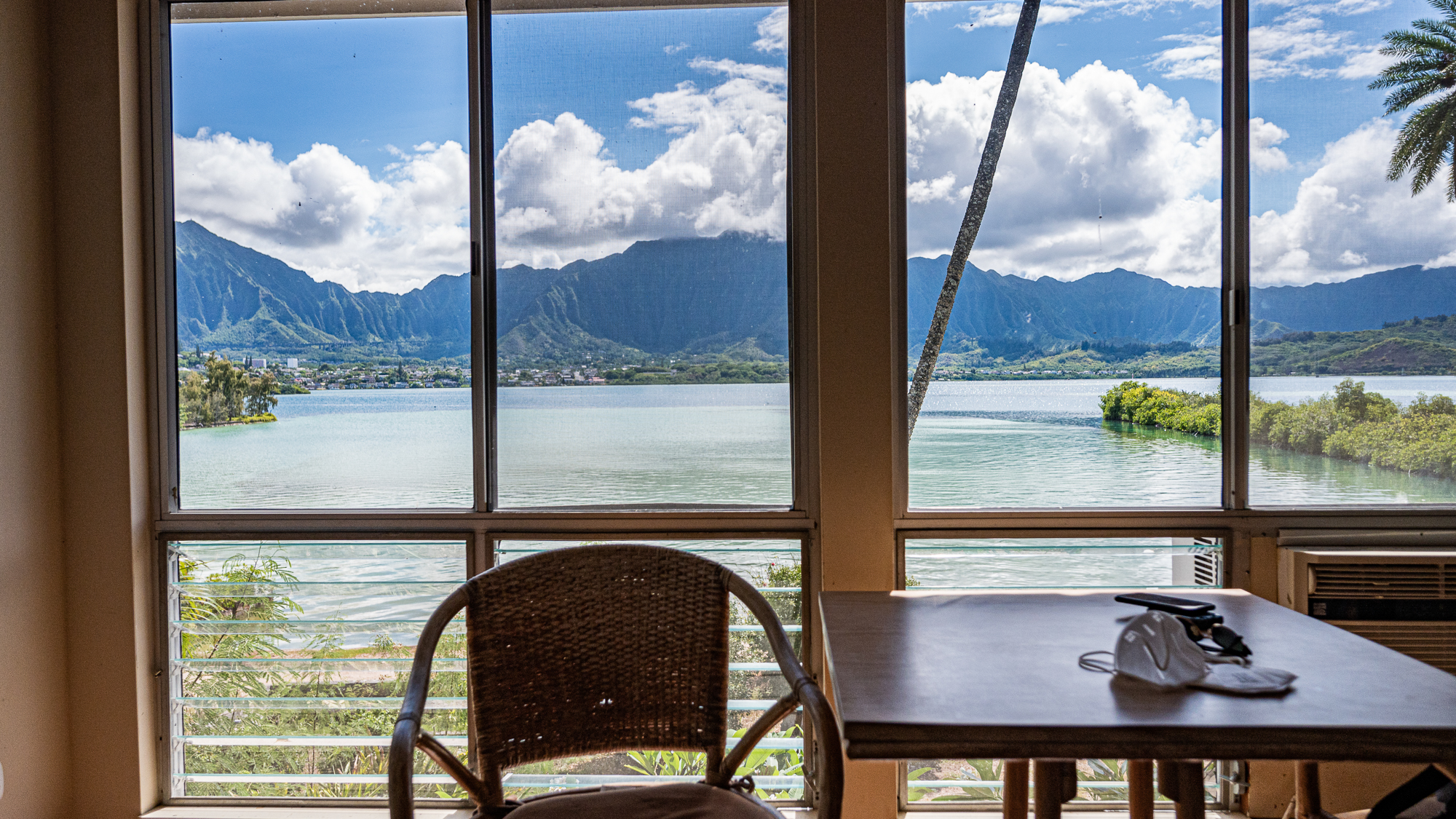
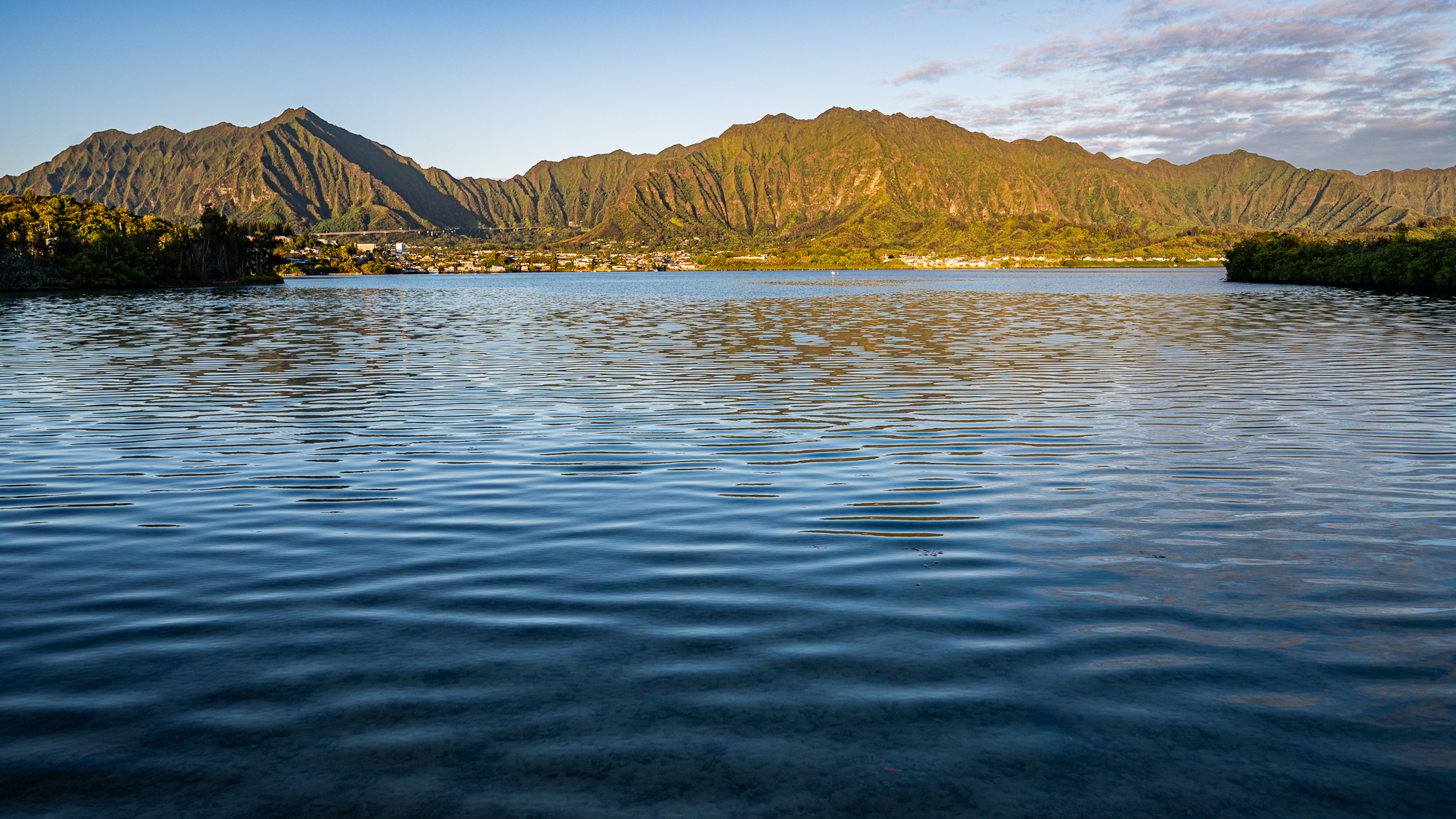
Among all the research institutions I have visited, HIMB takes the cake when it comes to scenic housing.
If you are a visiting scientist at the HIMB, you have a better view than anyone on the island of O’ahu. While everyone is looking at you and the vast ocean behind you, you are looking at the spectacular mountains of the island.
Well. After a very long journey we were finally there with our Zarges box, and it was actually a pretty good start.
August 17
As the first rays of Sun that would have passed our planet by if it wasn’t for the remnants of the ancient caldera were waking up my gecko roommates, my acid reflux was already ramping up its game: today was our last chance to cross our t’s and dot our i’s before the day of sampling.
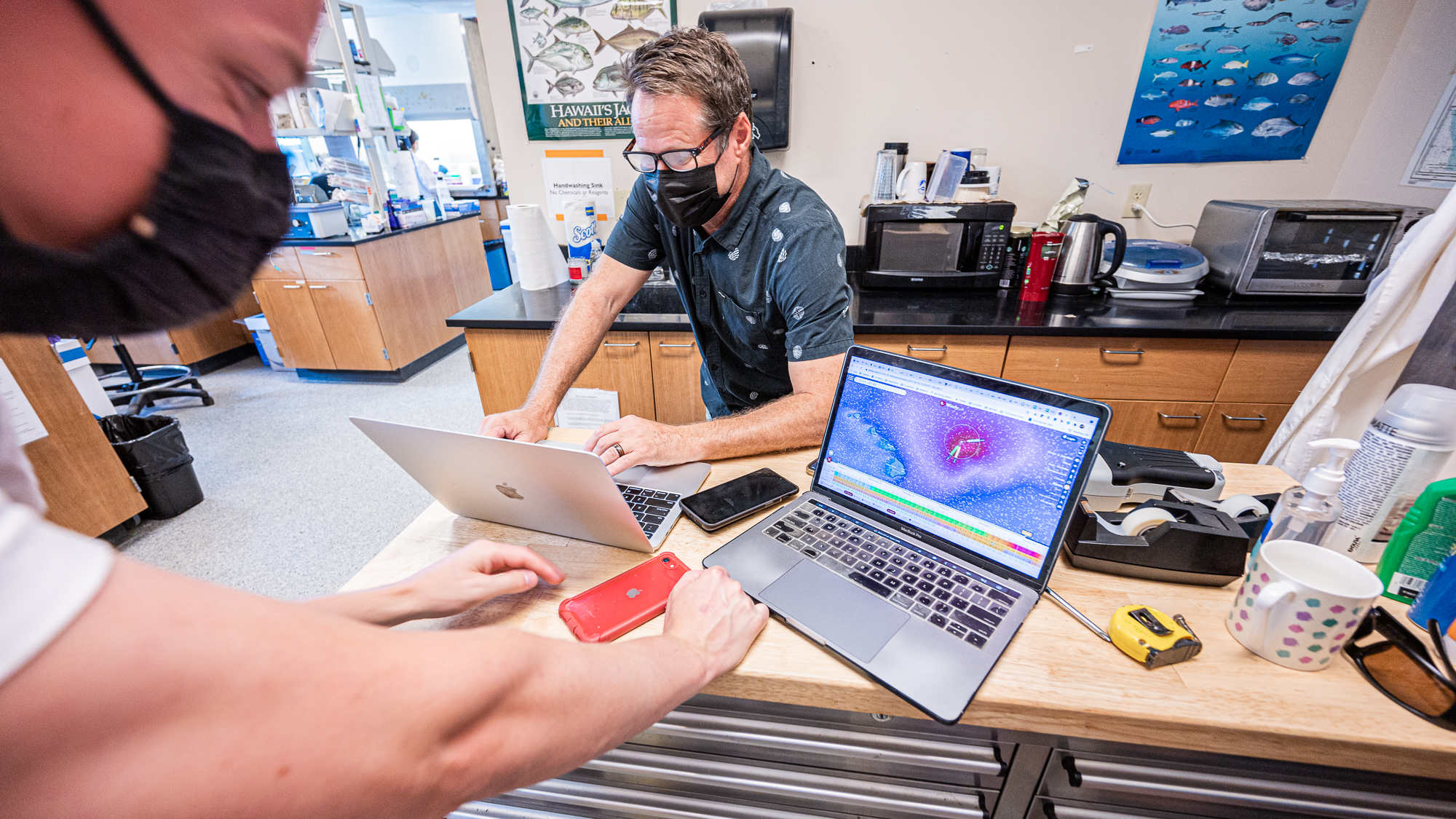
We walked from our housing to the Rappé lab headquarters. It was time to go through the plan as a group.
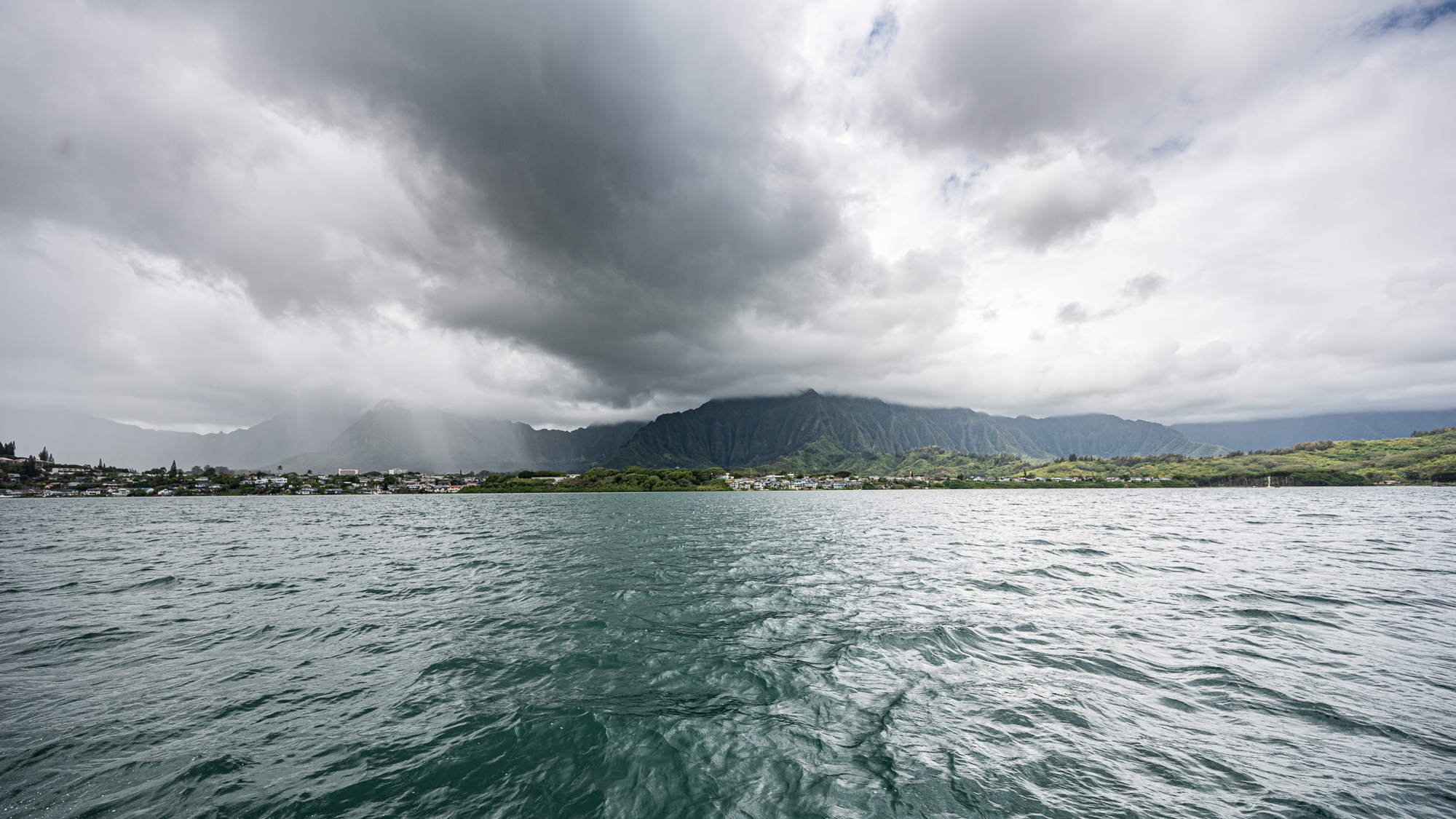
The first surprise was the fact that there was actually an incoming storm. My acid reflux was laughing like a maniac. It needed more than two days to arrive, but we realized that the swell for the second day during the night could be too dangerous to go out to the station outside of the bay.
We decided to move on with the sampling overall, and perhaps not collect samples from the open ocean station if the boat driver felt that the safety of the boat crew was anything less than 100%. Hearing all these conversations, my acid reflux was coming up to ask whether someone said we were not going to collect all samples. My acid reflux didn’t care about safety. It cared about getting the job done at any cost and no matter what. Luckily, Evan had some extra aluminum hydroxide & magnesium carbonate tablets for me to try.
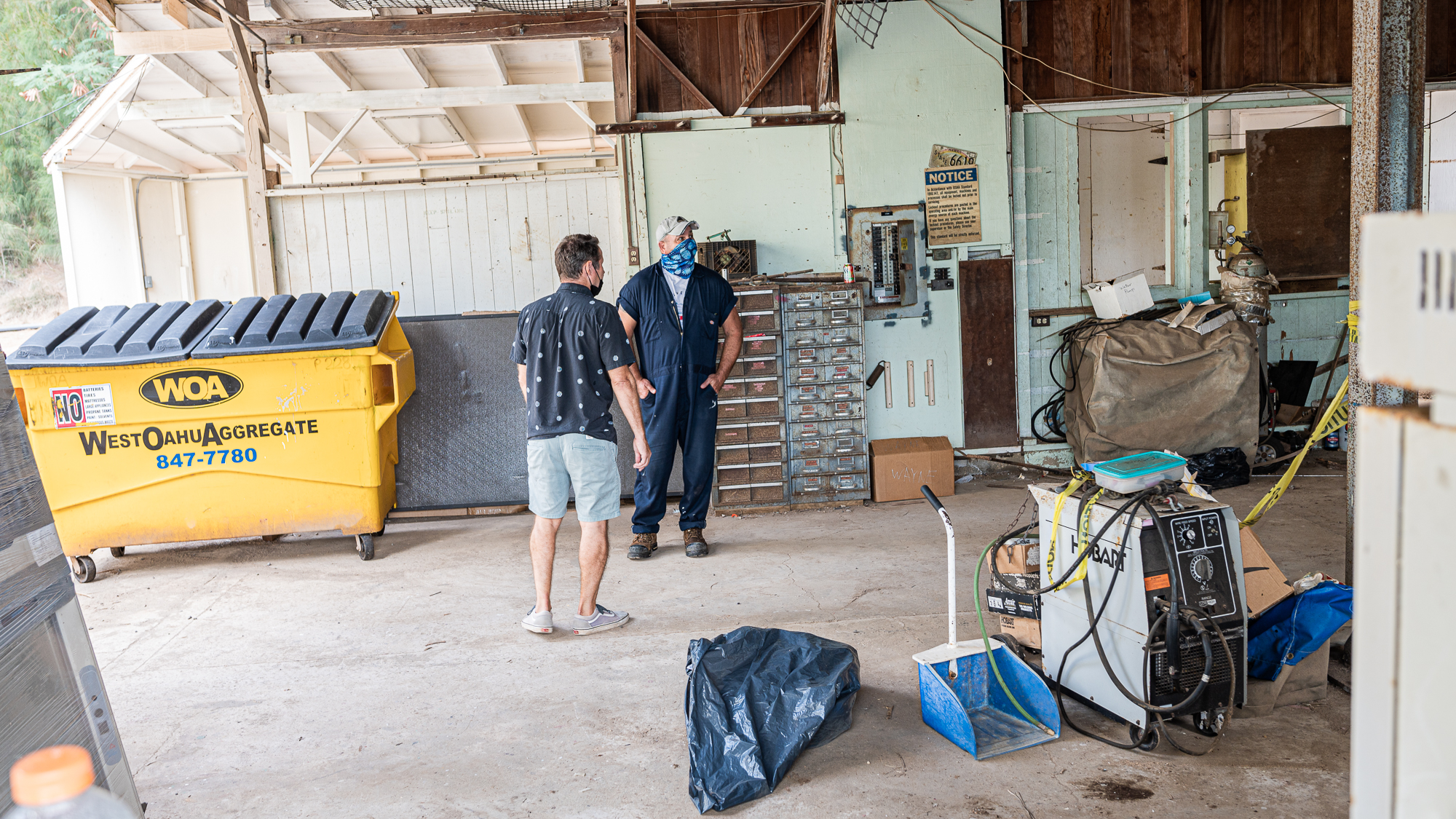
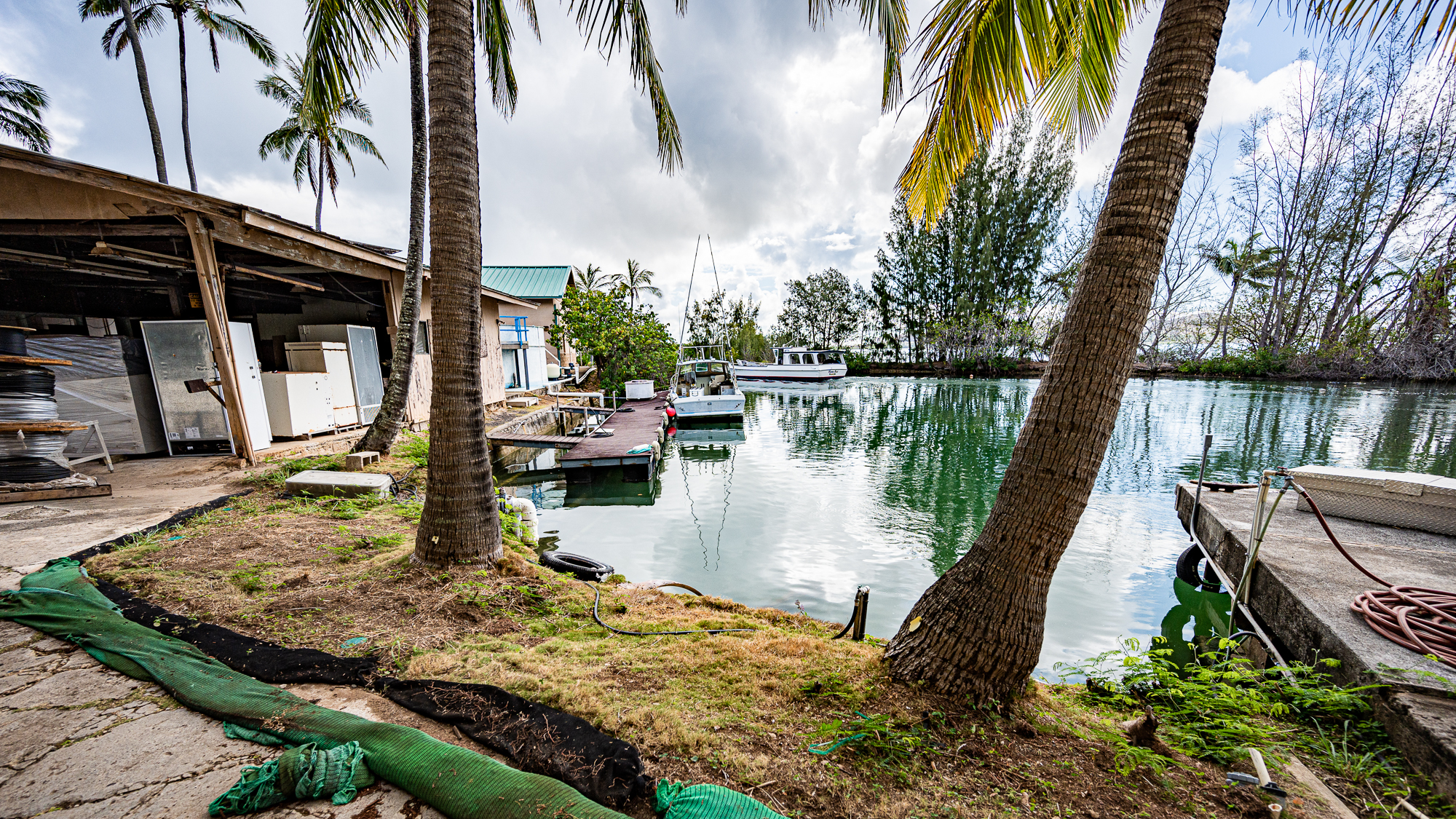
The next consideration was the lab space. During this continuous sampling effort, every full trip of the boat was going to take less than 90 minutes, leaving about 60 minutes for filtering for each batch of seawater. We were planning to carry carboys filled with water to Mike’s regular lab space for filtering, which was going to be quite a hike given the large number of heavy carboys we were going to bring. But Mike thought that we could actually put together a lab outside the building, right next to the dock.
It was going to take some effort to set it up, but then we will have a space only a few meters away from the dock, significantly cutting the transit time of carboys before filtration (the place in the photo above is the open building that can be seein the photo below).
I had always thought HIMB was a great place to do this lightweight but intense sampling expedition, it turned out to be much better than I could imagine.
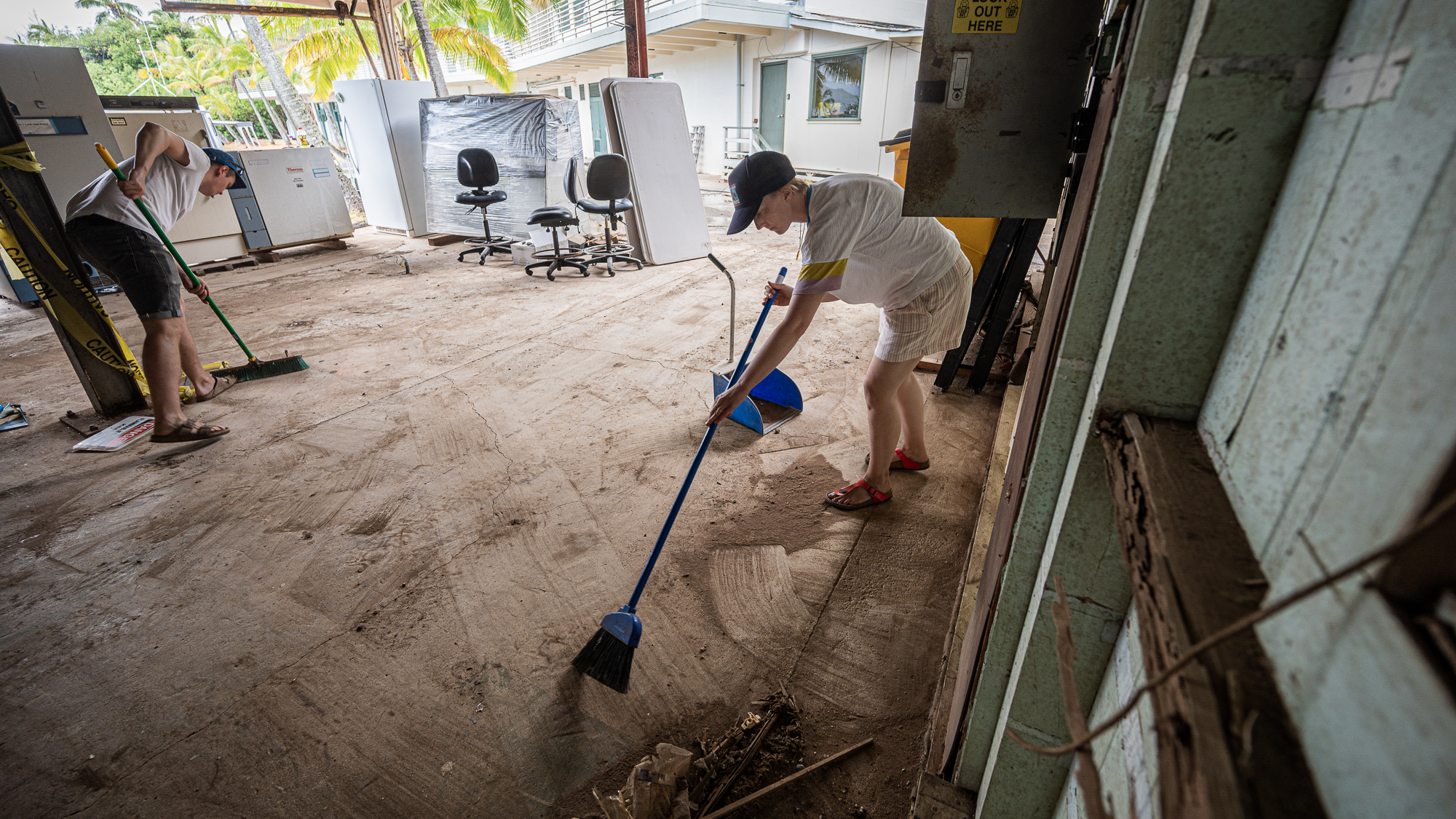
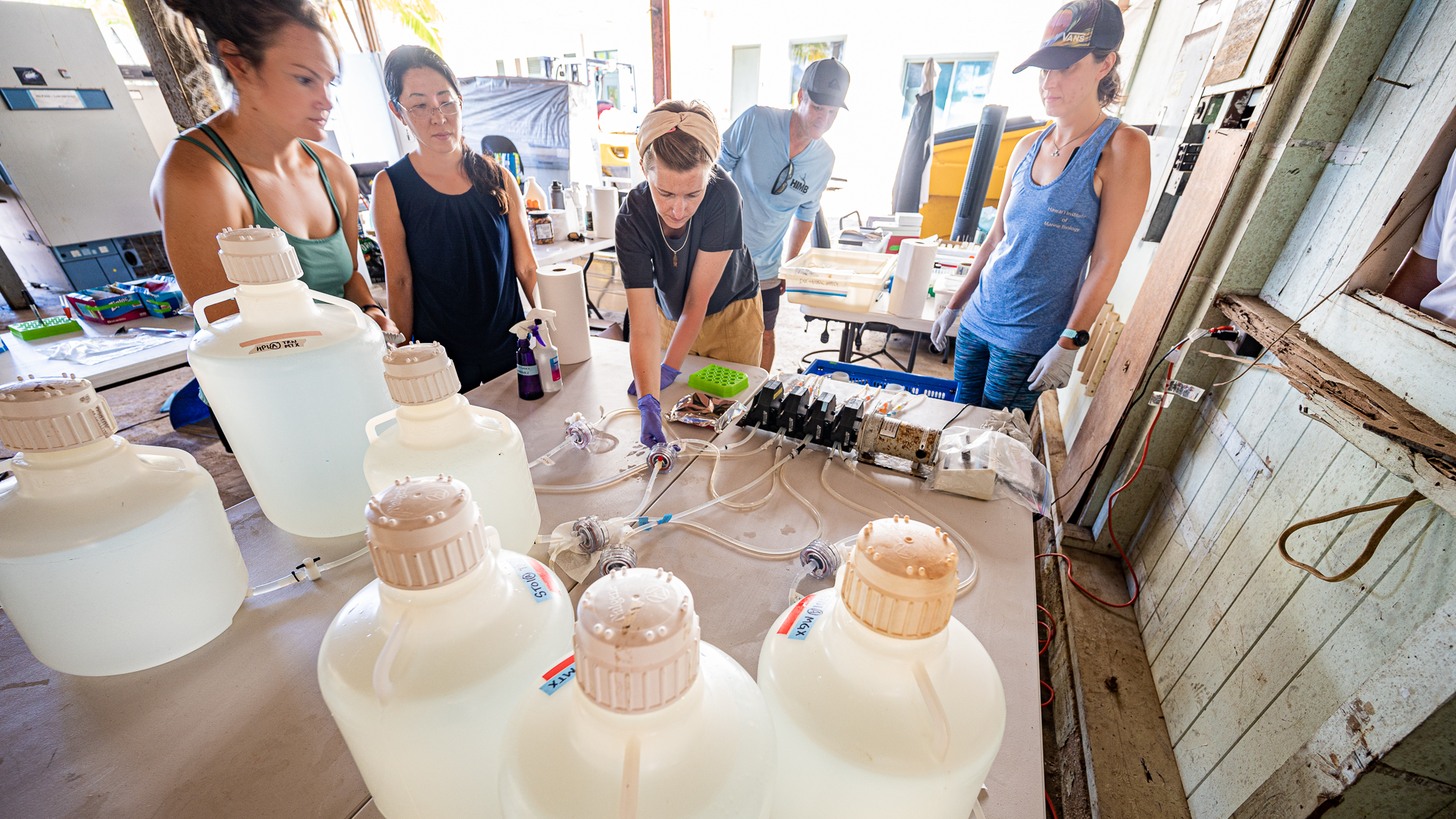
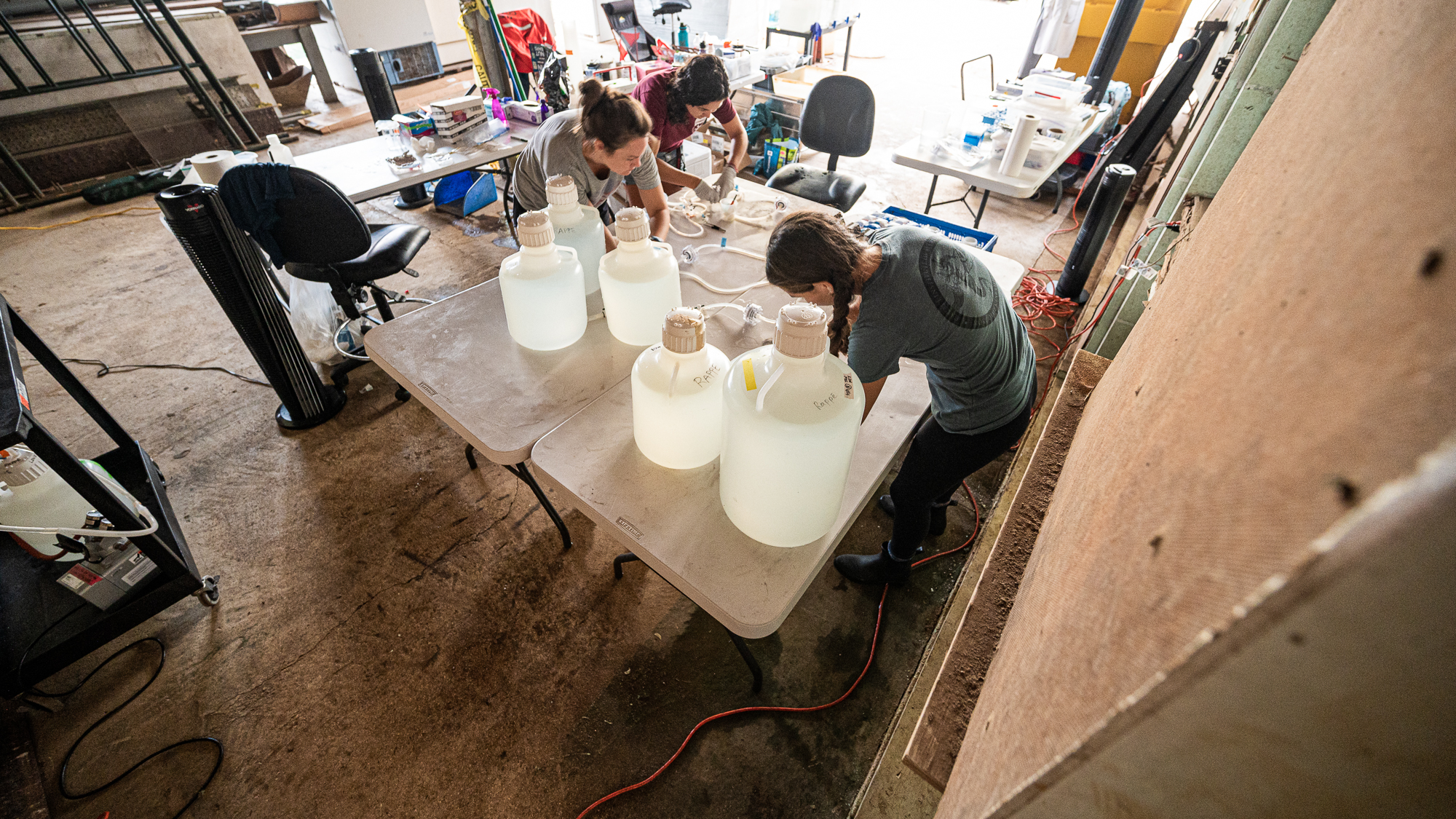
We first cleaned up the entire space,
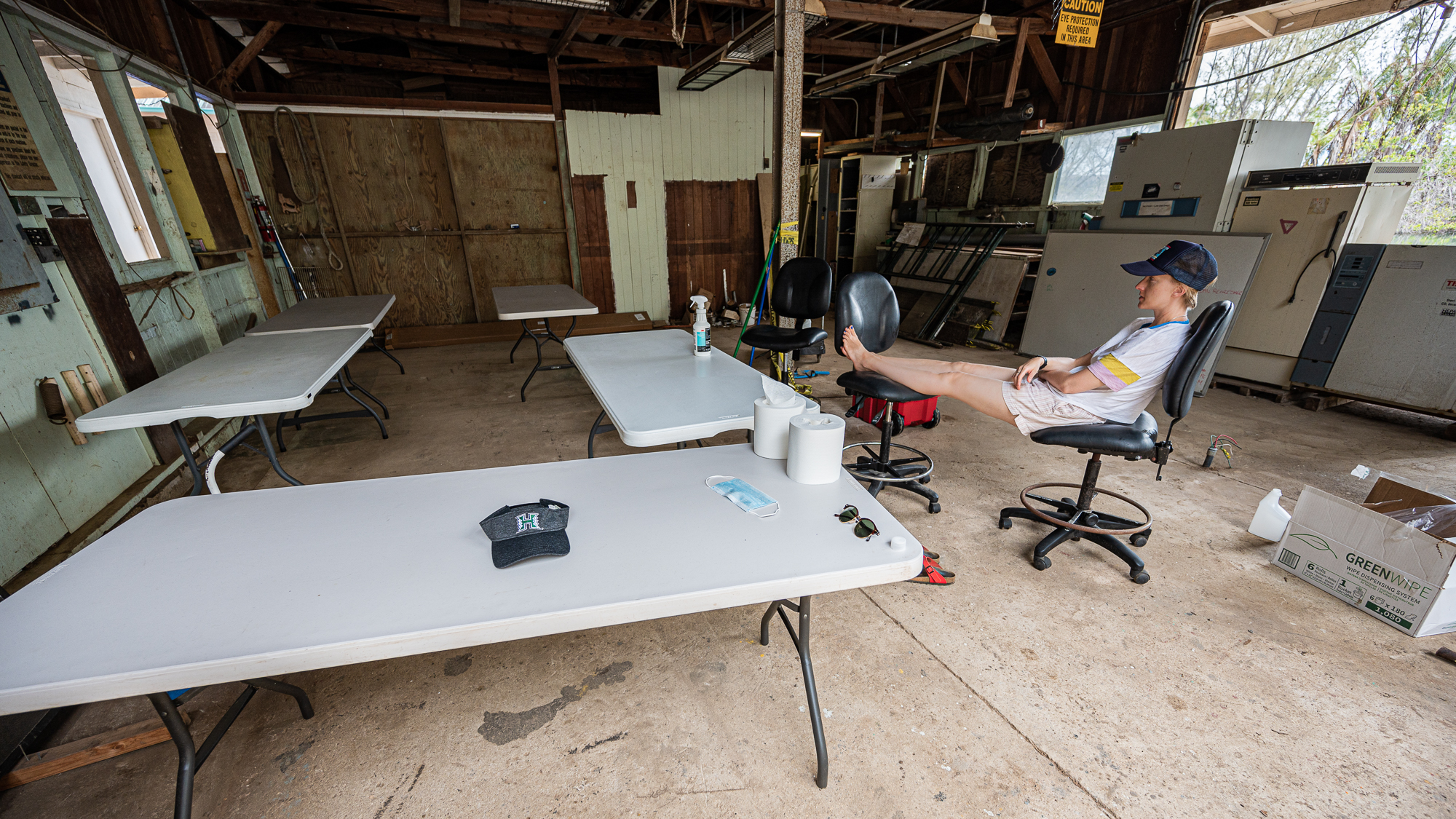
And then brought in some tables:
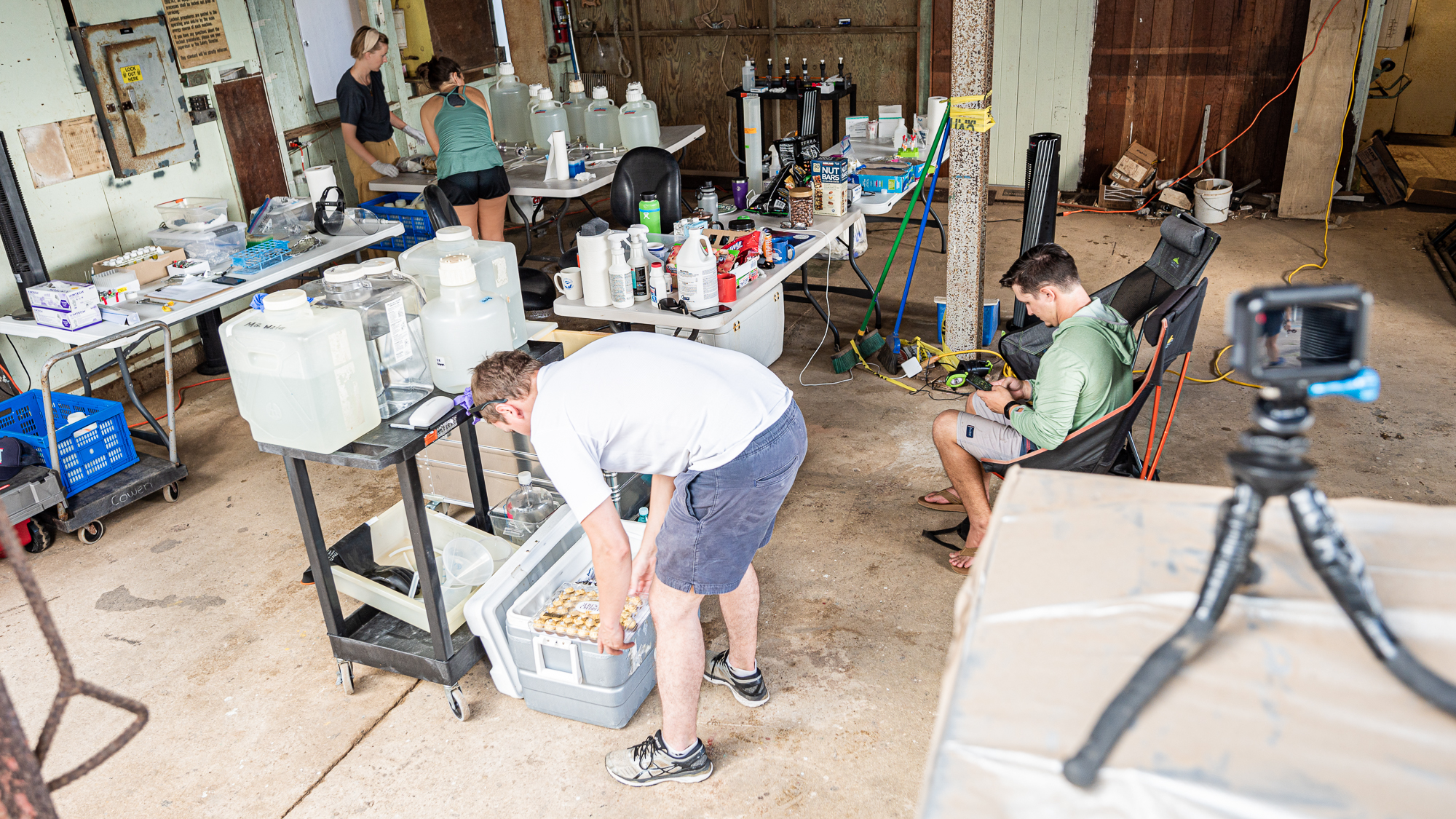
In just a few hours, we had a fully operational lab with every essential equipment for the completion of this project, including the 48 individually wrapped Ferrero Rocher hazelnut chocolates. Just right next to the dock. I think this was one of the most brilliant spontaneous decisions we’ve made during the sampling.
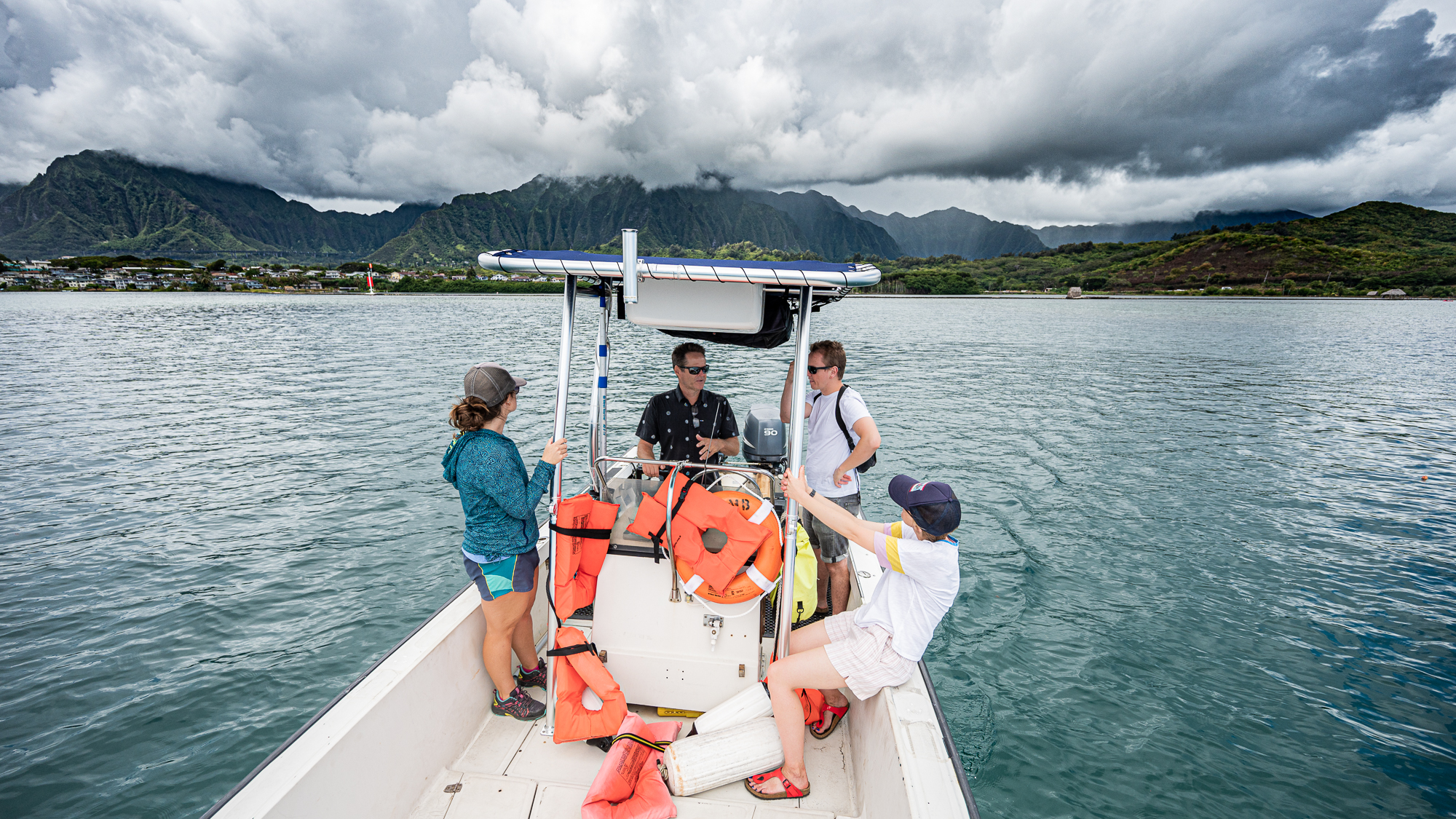
One of the last of things in our checklist for August 17 was to take a quick trip to the two stations, HP1 (inside the Kāne’ohe Bay, representing a coastal system) and STO1 (outside of the Kāne‘ohe Bay, representing an open ocean system), to have an idea about the distance between them, ideal path to traverse both, overall time a full round takes, and so on.
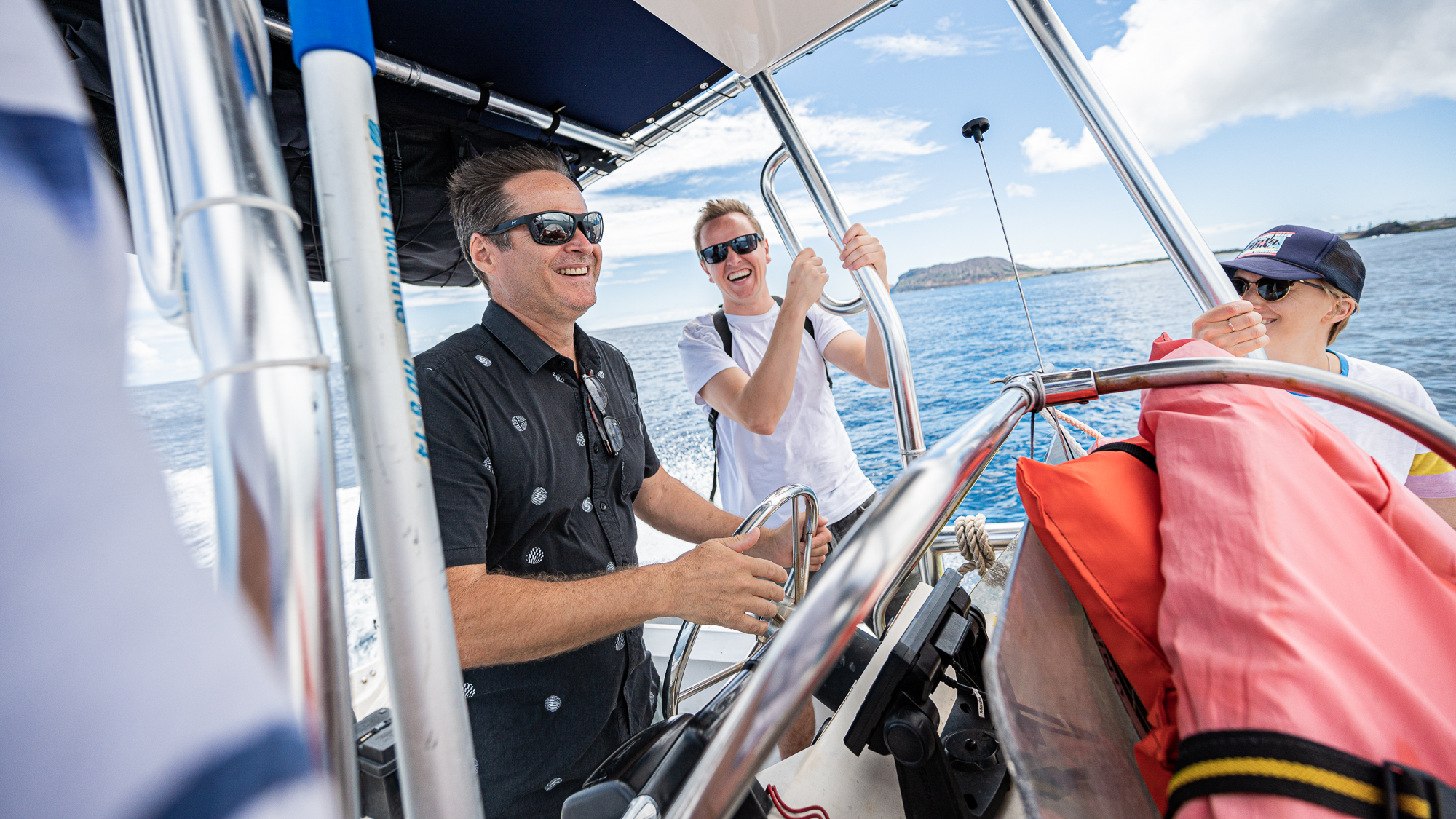
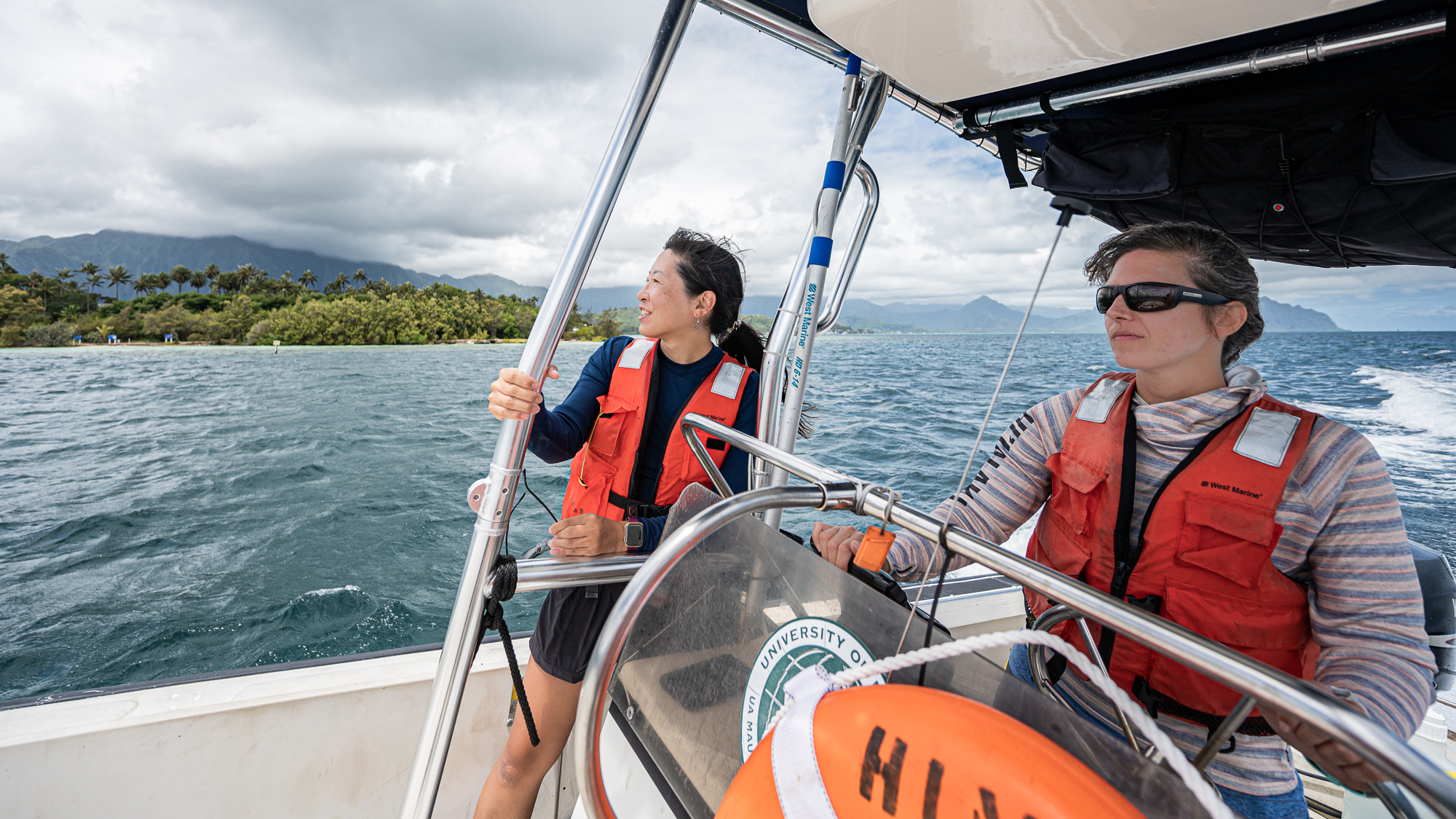
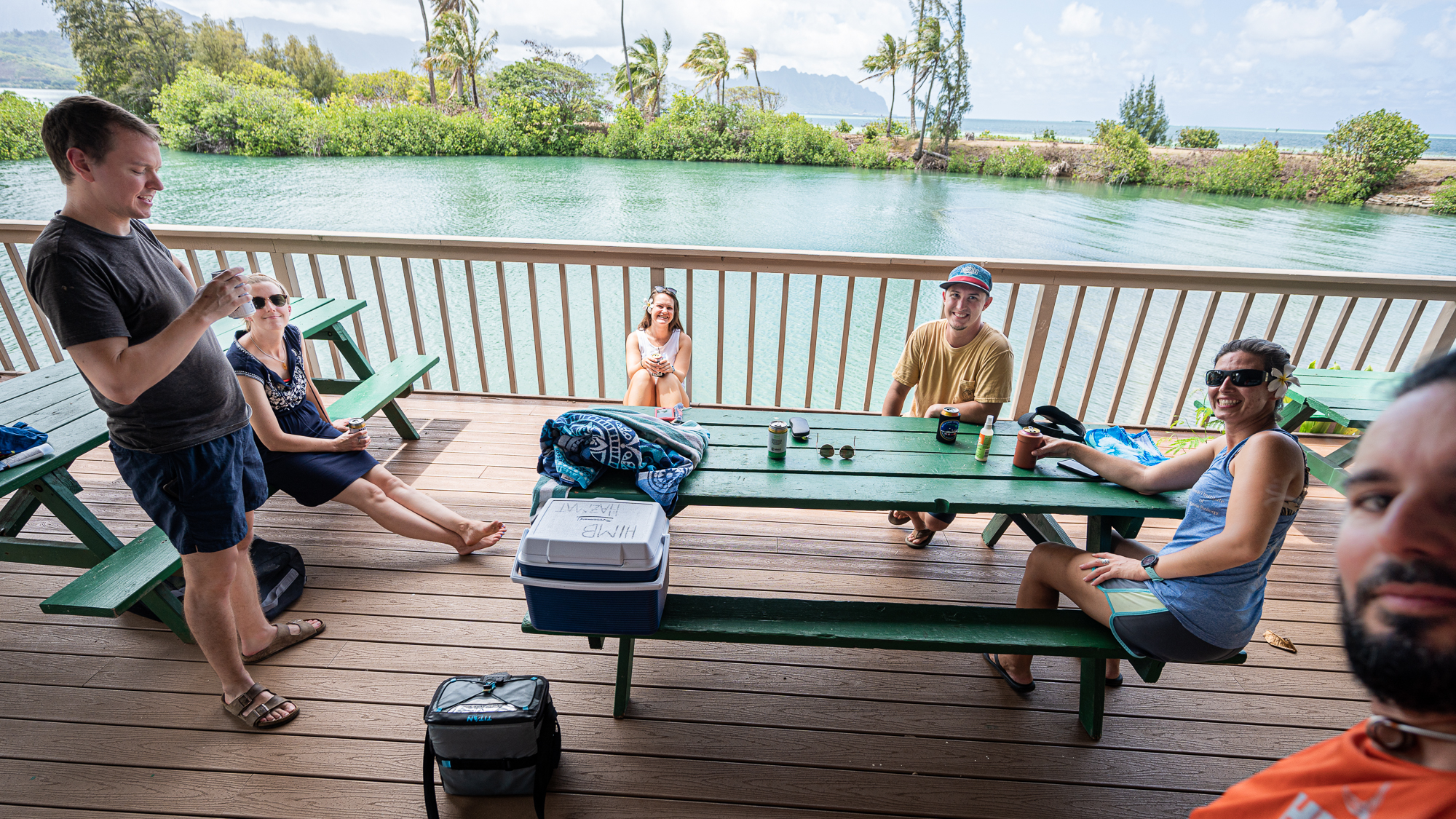
Things were looking good. Everyone was as happy as those people who appear in sunglass advertisements.
I wasn’t happy, though. I had to deal with my acid reflux and its stupid narration over every single thing “Hey. Yo. Wow. Are you guys seeing those clouds and rain and stuff? Wow. MUCH CLOUDS. Yup. We. Are. Doomed. We. Dead. Kaput. We better go home now. WHILE WE CAN. Srsly. SORRY, SIMONS FOUNDATION. MEREN HAS FAILED YOU. HE HAS FAILED EVERYONE :(”, etc.
No amount of aluminum hydroxide & magnesium carbonate could stop it, but the peaceful and quiet nights of Hawai’i helped.
But overall, my acid reflux that started a day before the trip did not stop until we were back in Chicago.
August 18
We were starting the sampling at 9am to follow a busy schedule:
These shifts included many more names than only Meren, Mike, Evan Kiefl, Jessika, Kelle, and Sarah. That is how this otherwise impossible sampling expedition became a reality thanks to those who volunteered their time, including,
-
Evan Barba, a graduate student at the Marine Biology Grad Program at the University of Hawai’i at Mānoa working on metazoan genomics with Rob Toonen in the Toonen-Bowen (ToBo) Lab. Evan’s research focuses on population genomics of invertebrates, fish, and corals, with a goal of informing management and conservation efforts. He also is a member of the ToBo lab’s bioinformatics development team, generating data processing pipelines. After his PhD, he hopes to work at the agency level - either government or NGO to conserve marine wildlife. Evan contributed to the sampling effort as one of the boat operators. And he also contributed the title of this blog post! When I told him that I was planning to document the entire journey of this project, he said “from boats to bits, then?”. Thank you, cap’n Barba!
-
Andrian Gajigan, who is better known as Adi, studies microbial oceanography, marine virology, genomics, and gene regulation as an Oceanography graduate student at the University of Hawai’i at Mānoa with the Steward Lab. After his PhD, he plans to continue doing research as a postdoc in the continental US and in Europe before finally applying for a faculty position in the Philippines or anywhere in Southeast Asia and the Pacific. Adi contributed to the sampling effort by helping both in processing samples in the lab and sampling in the boat.
-
Clarisse Sullivan, am Oceanography graduate student at the University of Hawai’i at Mānoa. She is interested in studying microorganisms that thrive in the thermodynamic limits of life. Clarisse contributed to the sampling effort by helping collect surface seawater during the evening to sunrise shifts (when everyone else was sleeping!).
-
Oscar Ramfelt, a graduate student at the University of Hawai’i at Mānoa, interested in both studying marine microbes and the application of computer programming to science. After his PhD, he plans to focus more heavily on programming and how it can be applied to advance science. Oscar contributed to the sampling effort by collecting seawater samples and processing them.
-
Mariana Rocha de Souza, a graduate student at the Coral Resilience and ToBo Labs at the HIMB, focusing on coral reefs and the role of the microscopic algae that lives in the coral in providing coral resilience to climate change. She is starting a Knauss fellowship on Marine Policy! Mariana contributed to the sampling effort by helping in the lab by collecting environmental data in the boat and by filtering water and in the lab.
-
Mariko Quinn, a sophomore at the University of Hawai’i at Mānoa majoring in Global Environmental Science, interested in coral reef ecology and reproduction. After the undergraduate education, Mariko plans to pursue graduate school for marine science. Mariko contributed to the sampling effort by filtering water samples and in the lab.
-
Ciara Ratum, an undergraduate student at Hawai’i Pacific University, who is interested in marine biology as well as communicating marine science especially to the local communities in Hawai’i. After the undergrad, she plans to work in a job related to marine biology before applying for graduate school. Ciara contributed to the sampling effort by collecting environmental data for the last three hours of the time series.
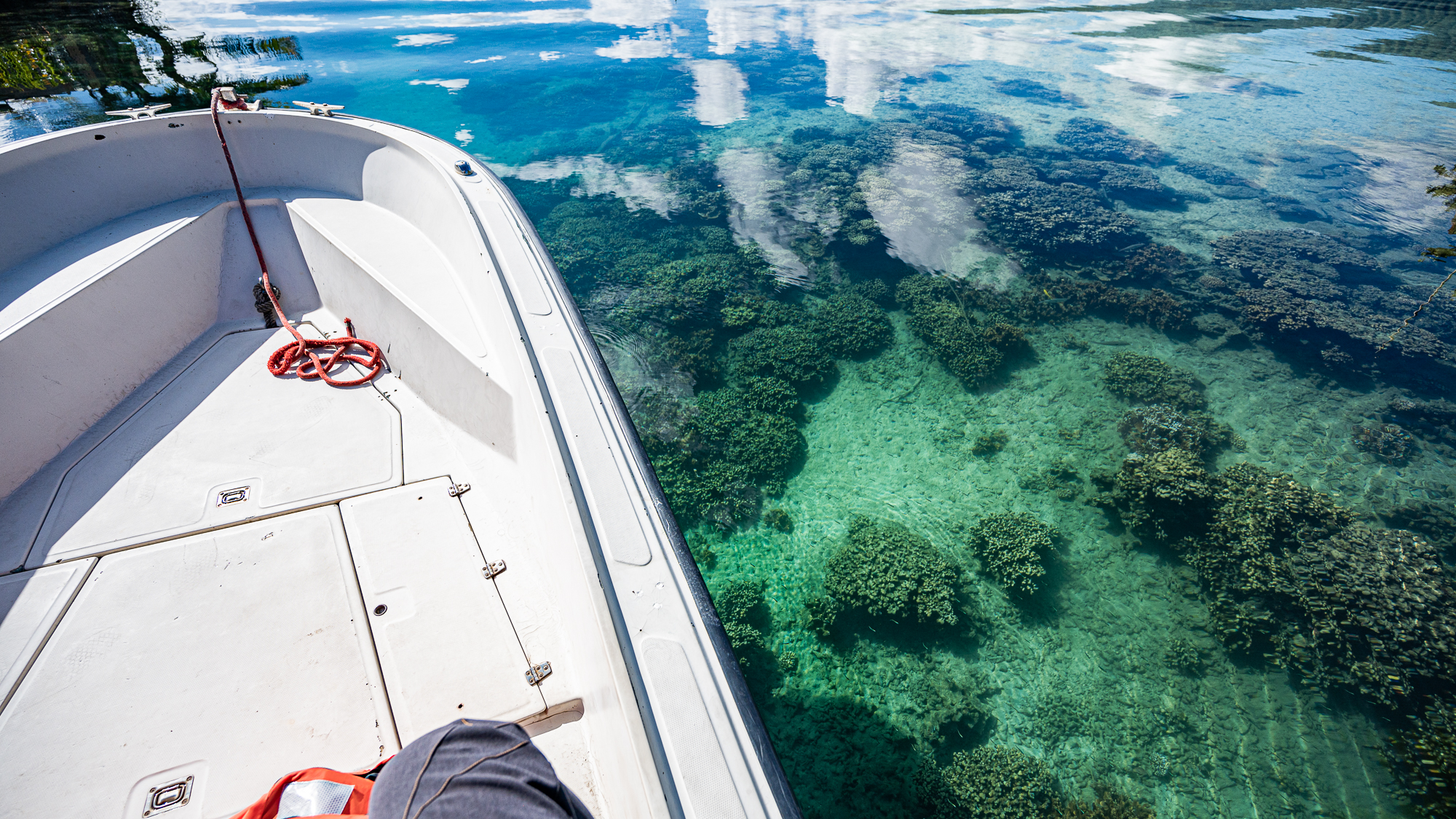
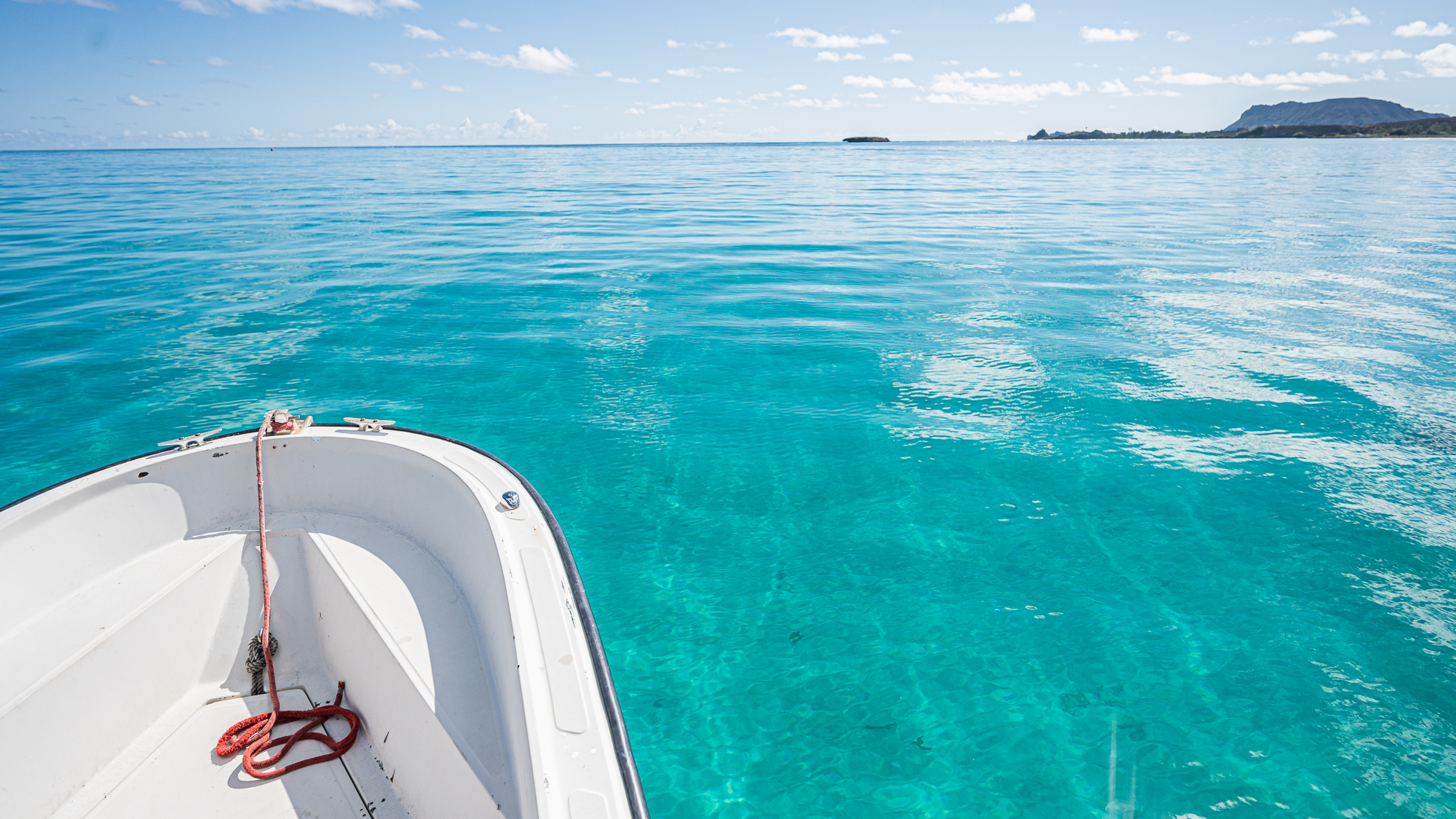
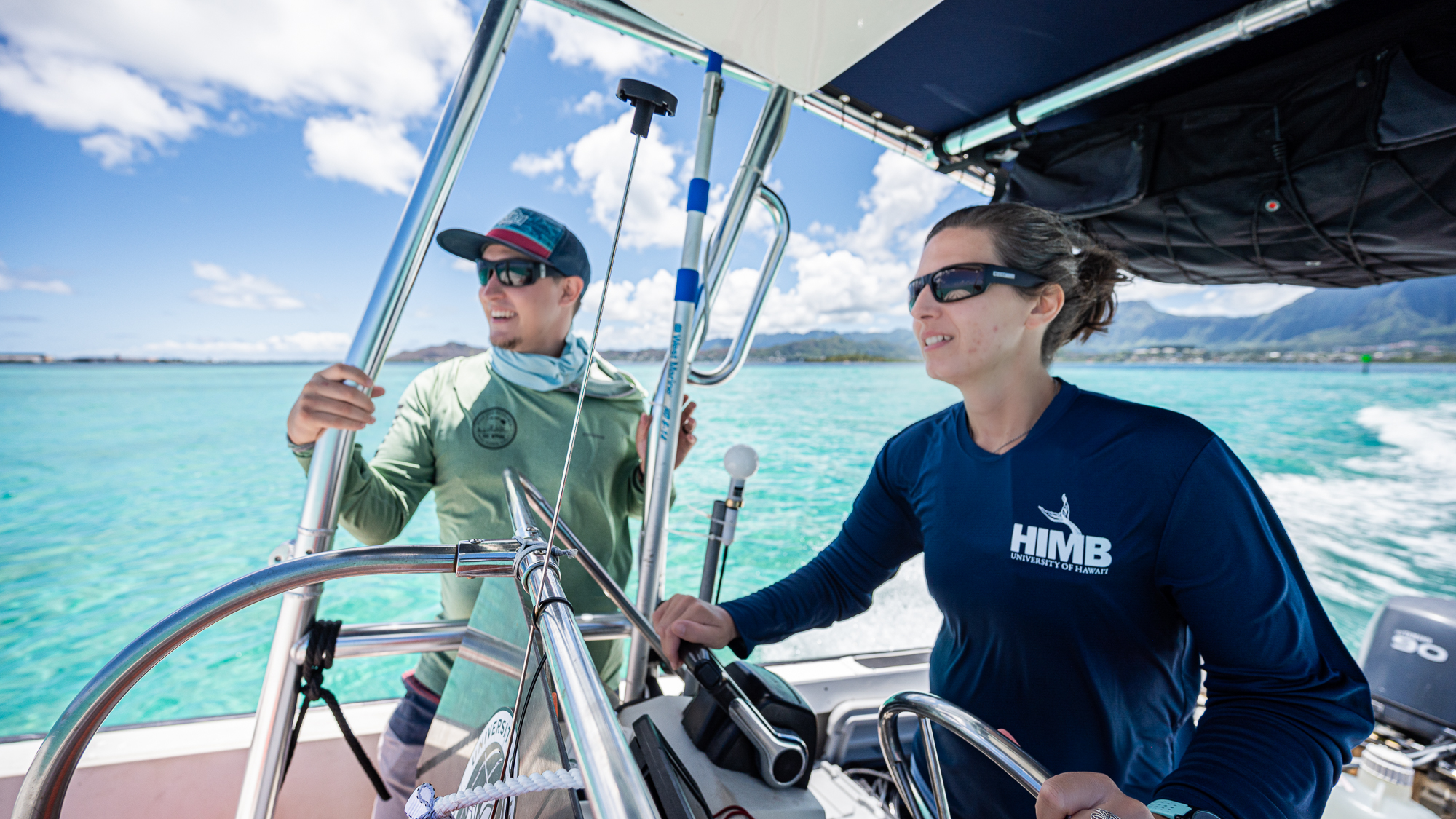
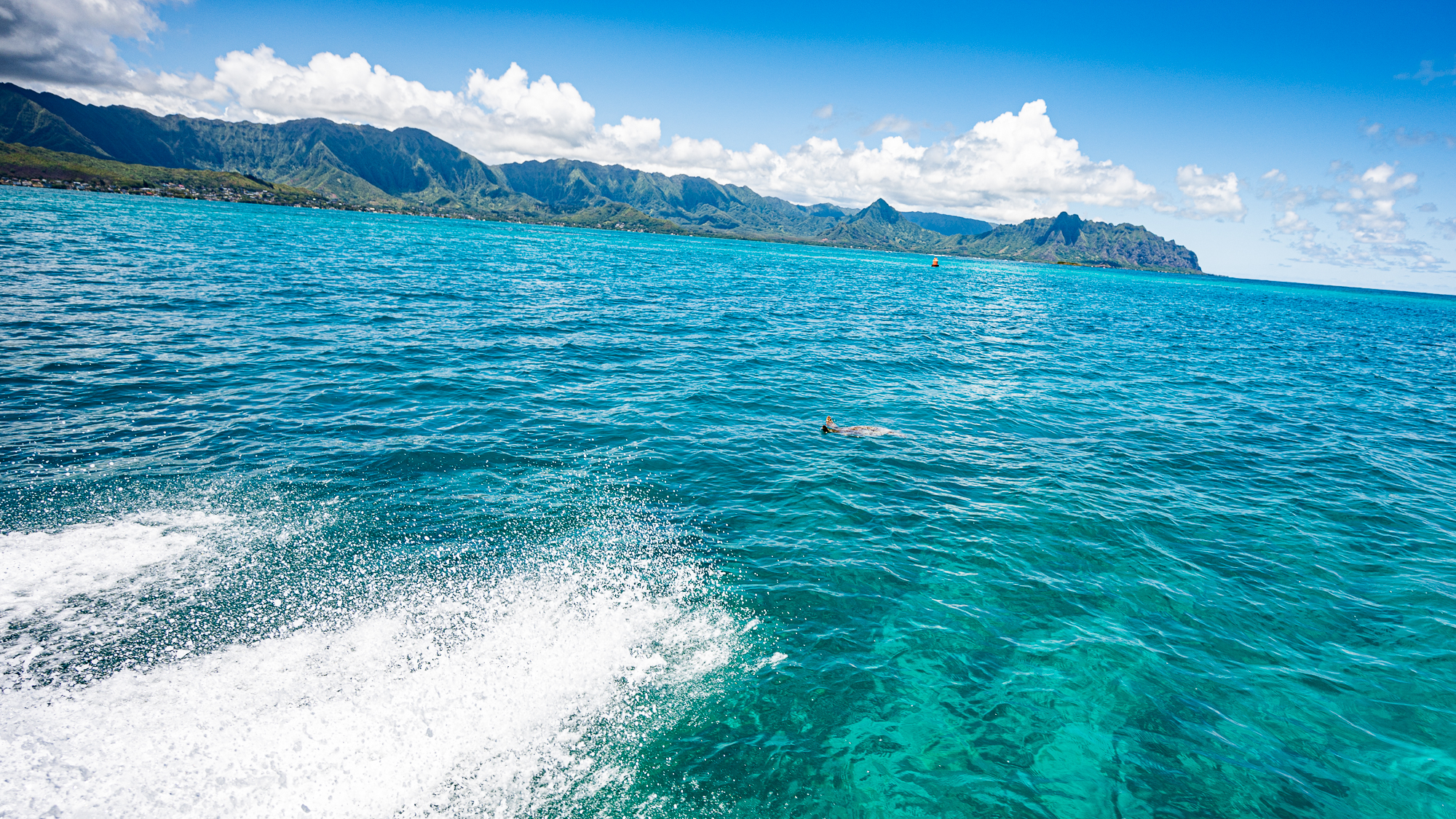
Despite all the scary clouds the day before, the water was crystal clear during our first trip at 9am.
Calling these waters ‘calm’ would have been injustice as they were almost completely flat with no waves.
So this first trip with our payload was going to be the fastest and smoothest of the entire sampling trip.
The legacy Mike Rappé left behind with that first trip was going to loom over the heads of our other captains, Kelle, and Evan Barba for the next two days :p
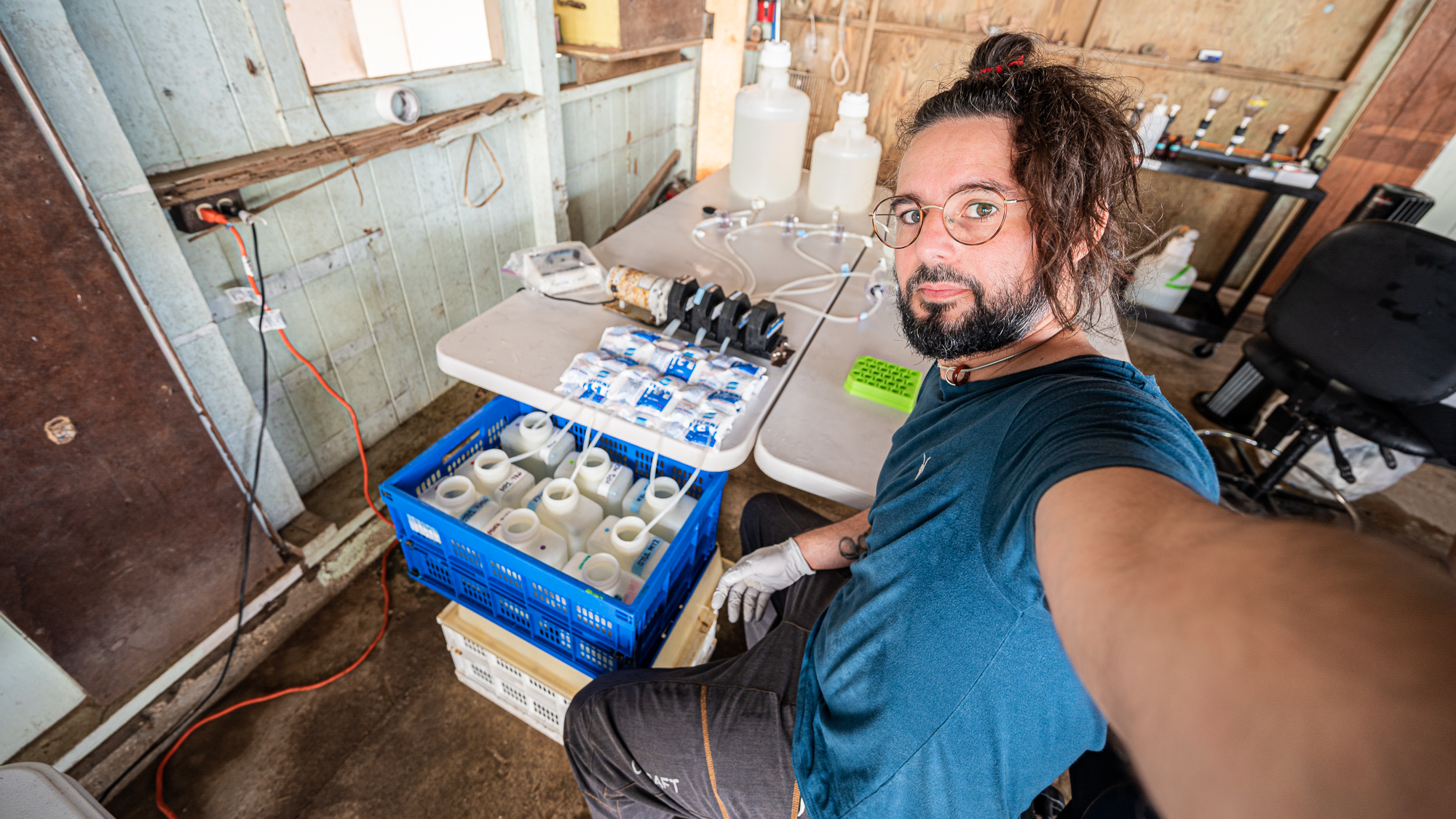
With our arrival to the lab, the very first round of samples were finally marking the beginning of filtering (under the supervision of Sarah, Hannah, Jessika, Mike, Kelle, and Evan),
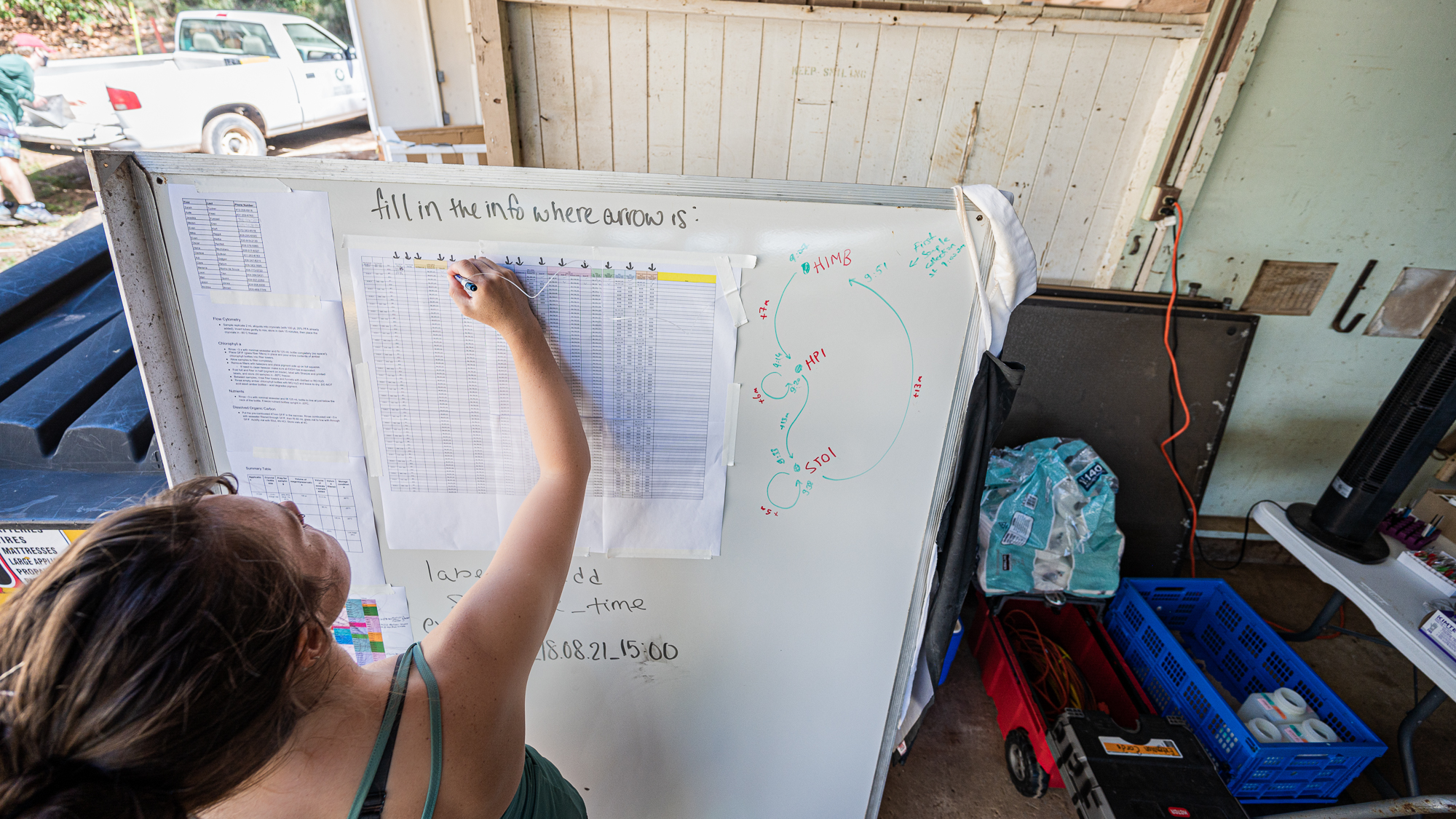
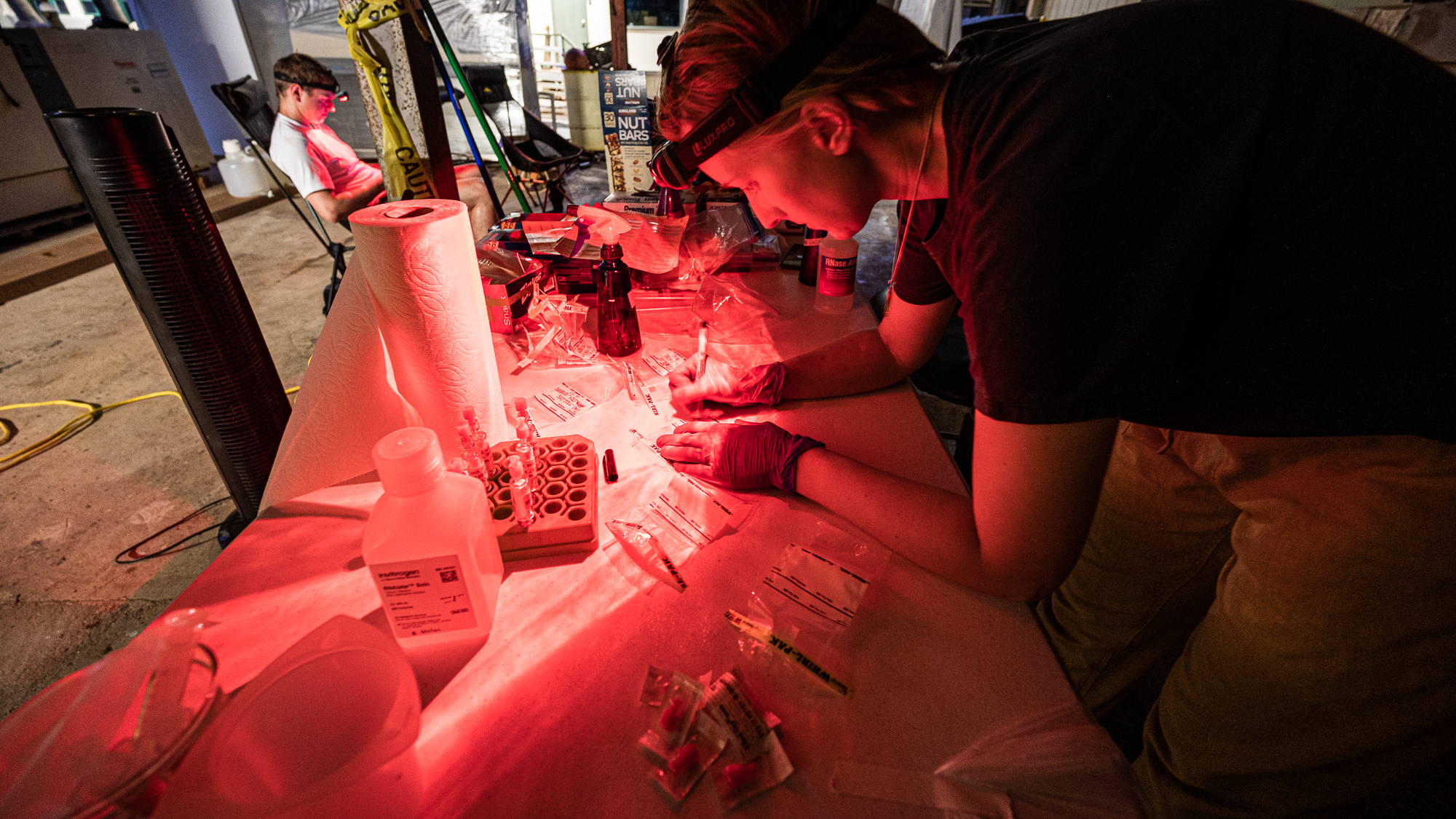
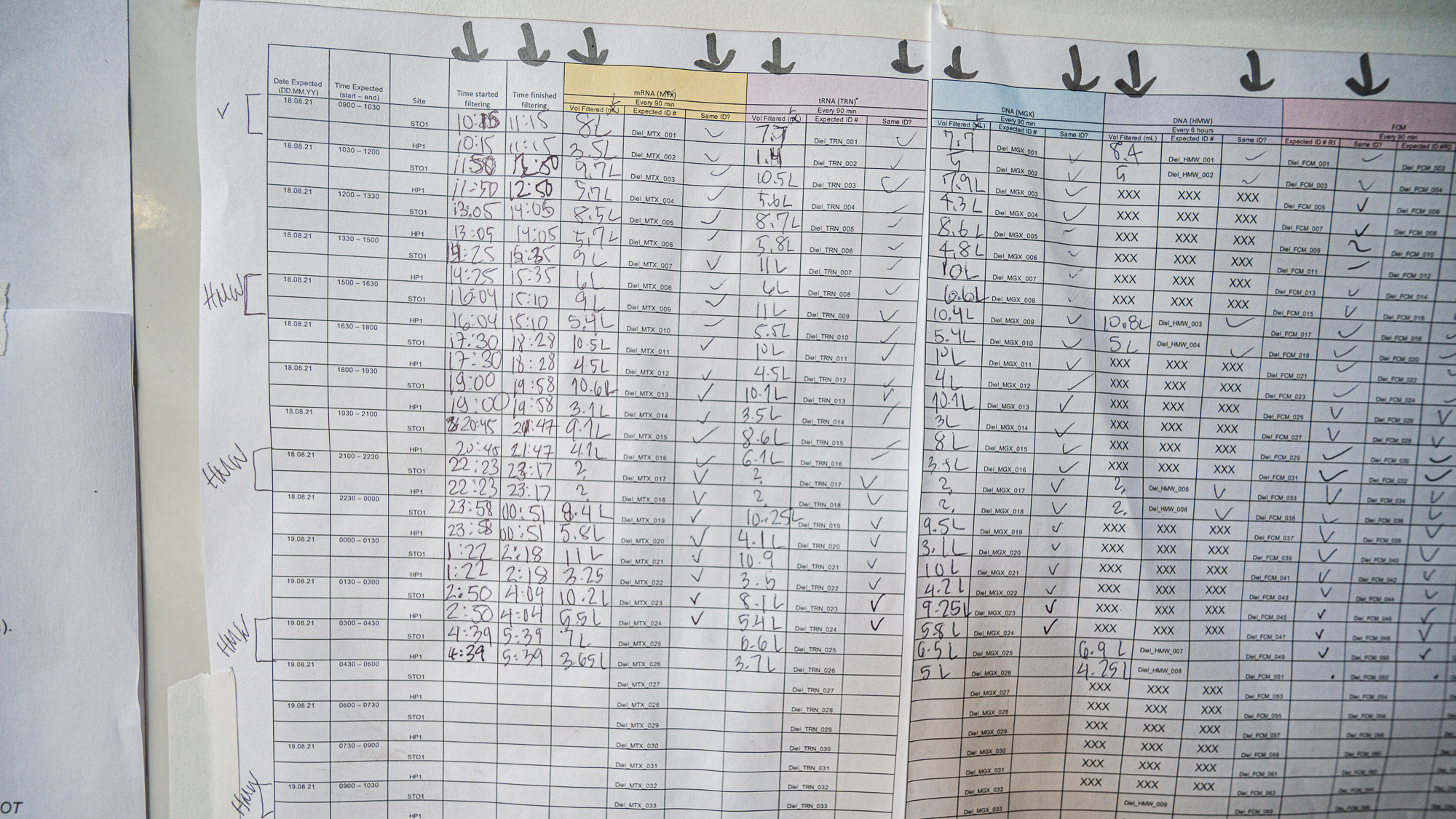
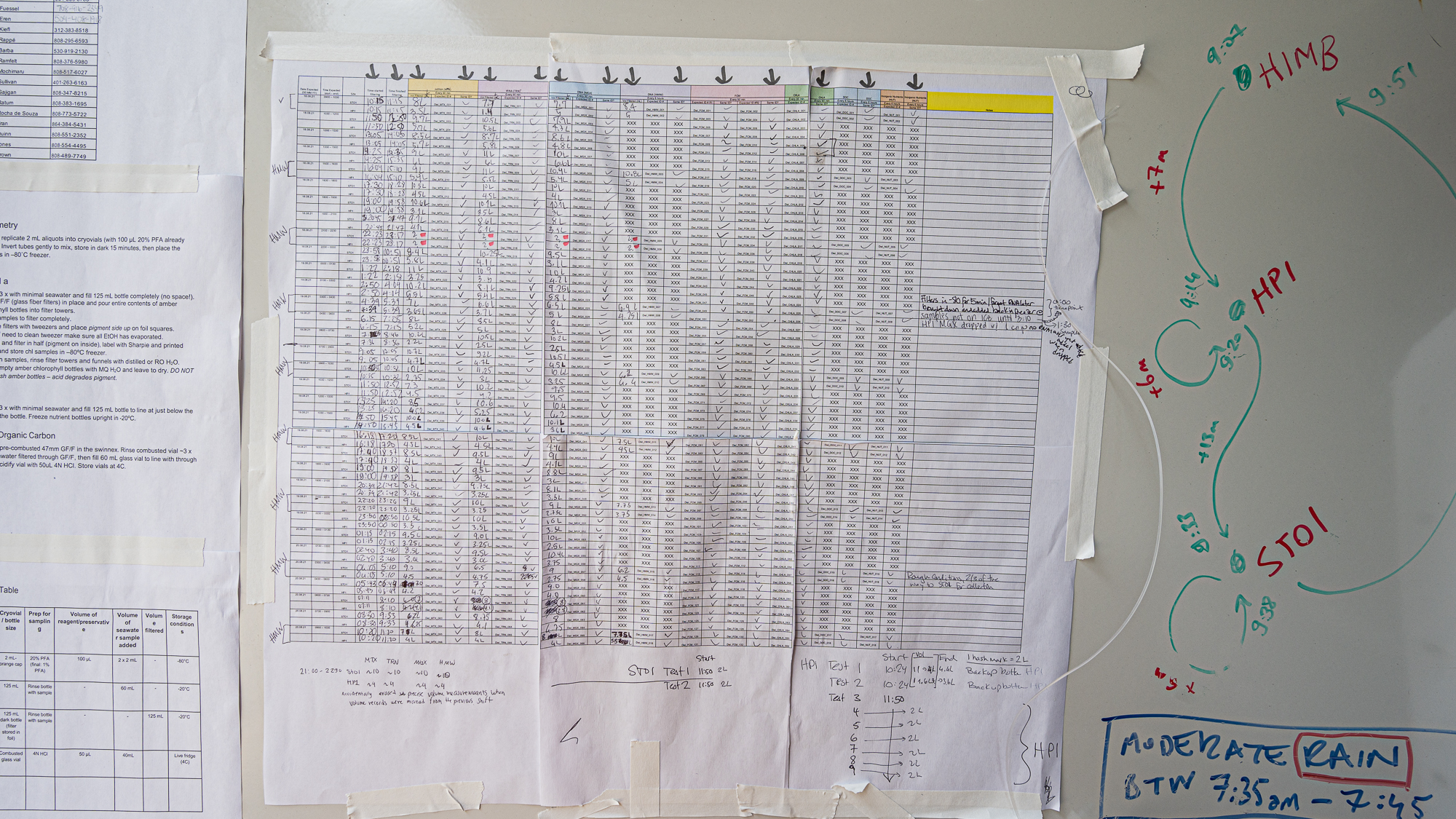
and moments later Sarah was putting in the very first entry of our long sample sheet with the timestamp for the beginning of the filtration of the first round of samples (while my acid reflux was whispering “lol there is no way you guys will be able to fill this up” – it had something to say about everything really).
While everyone was being very busy at the lab,
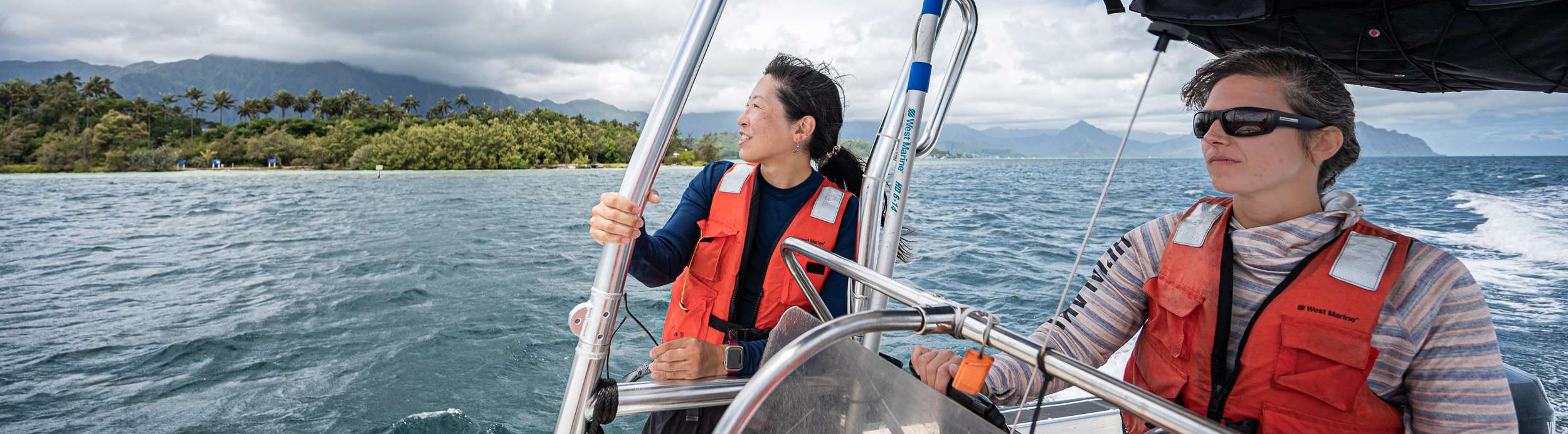
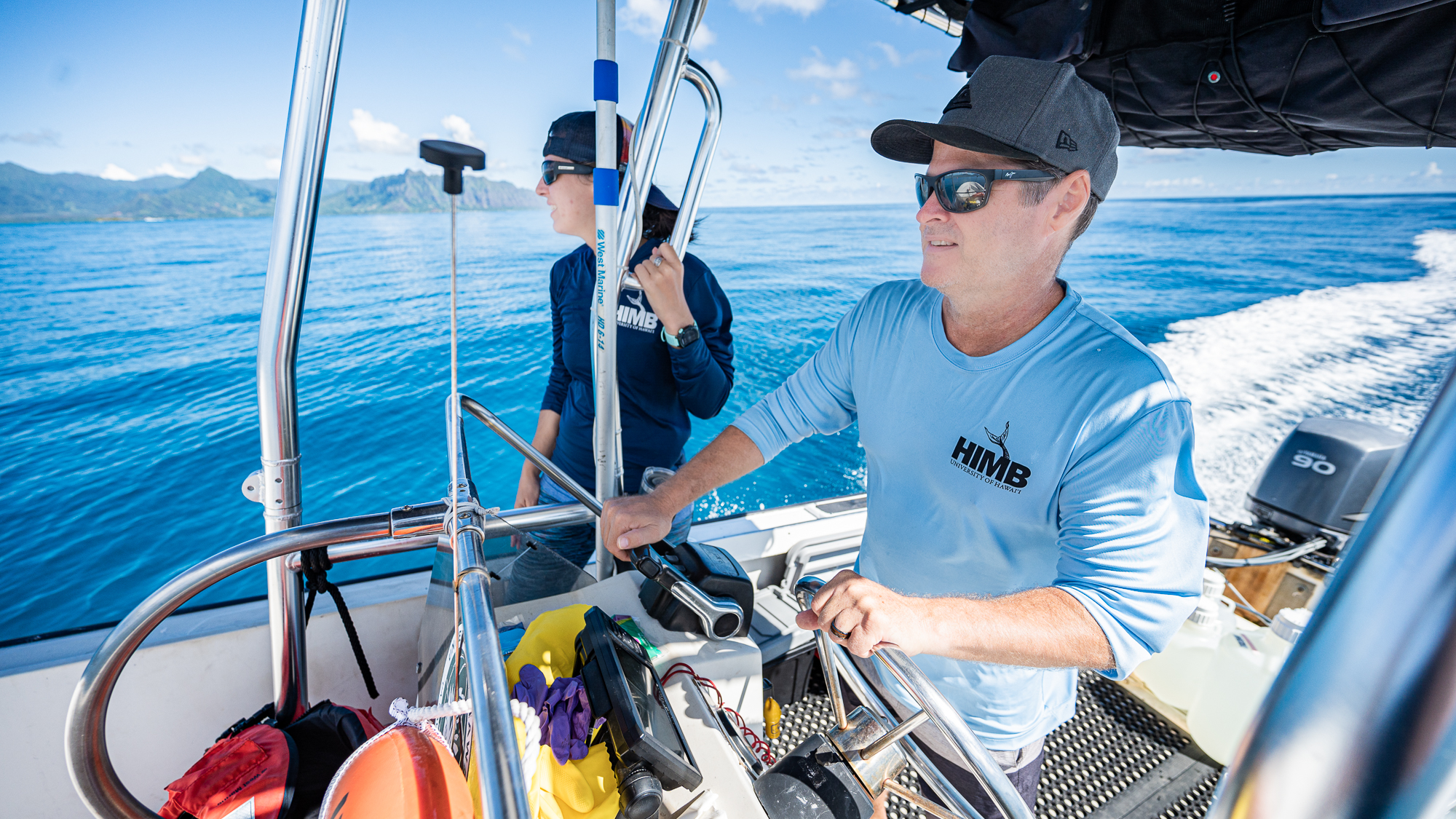
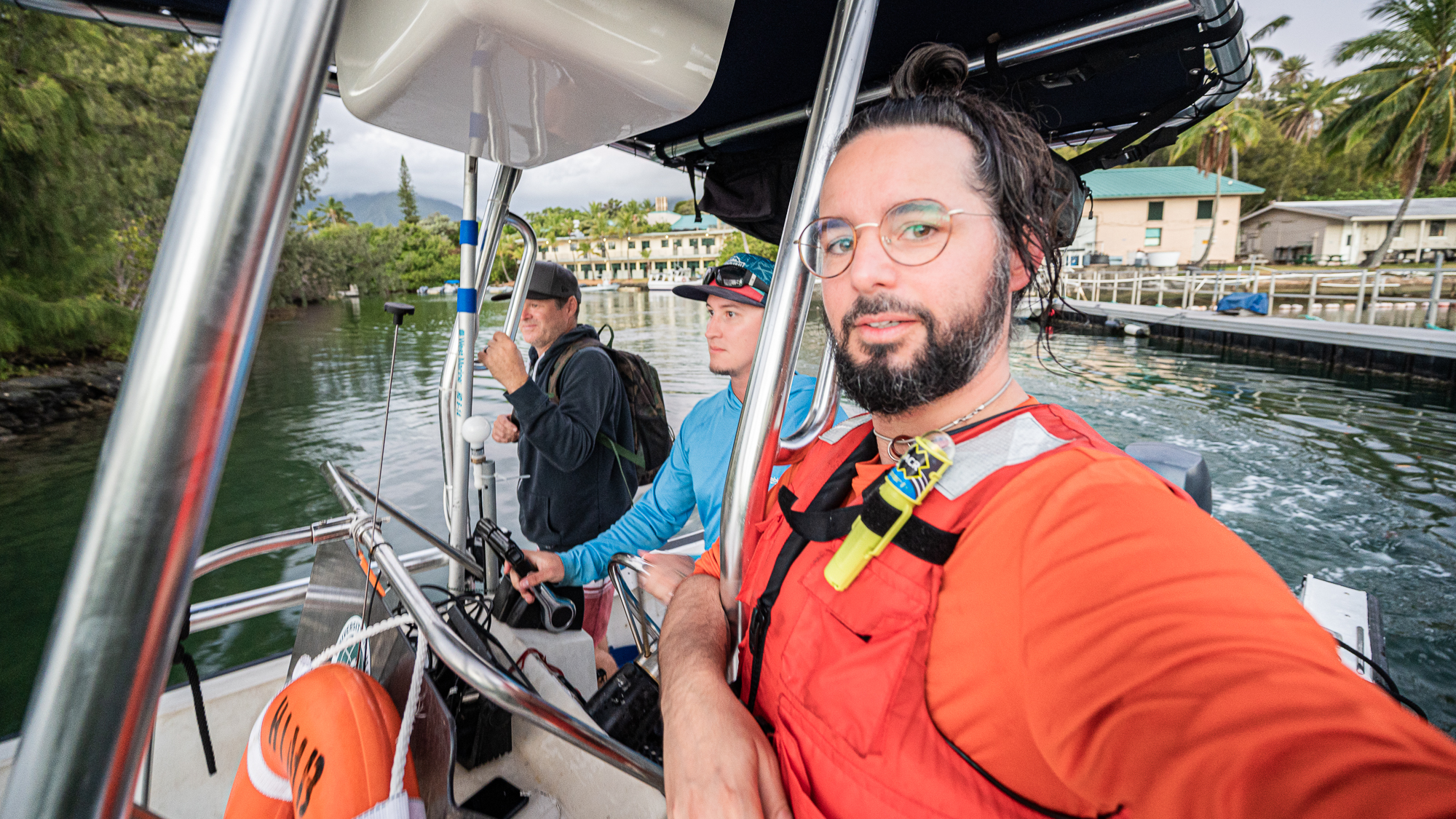
We, Evan Barba, Kelle, and myself, who were known to the rest of the grouop as the ‘sea people’ due to our inexplicable hardiness, were back on the ‘road’,
Being greeted by passing by sea turtles and such.
Everyone had a job.
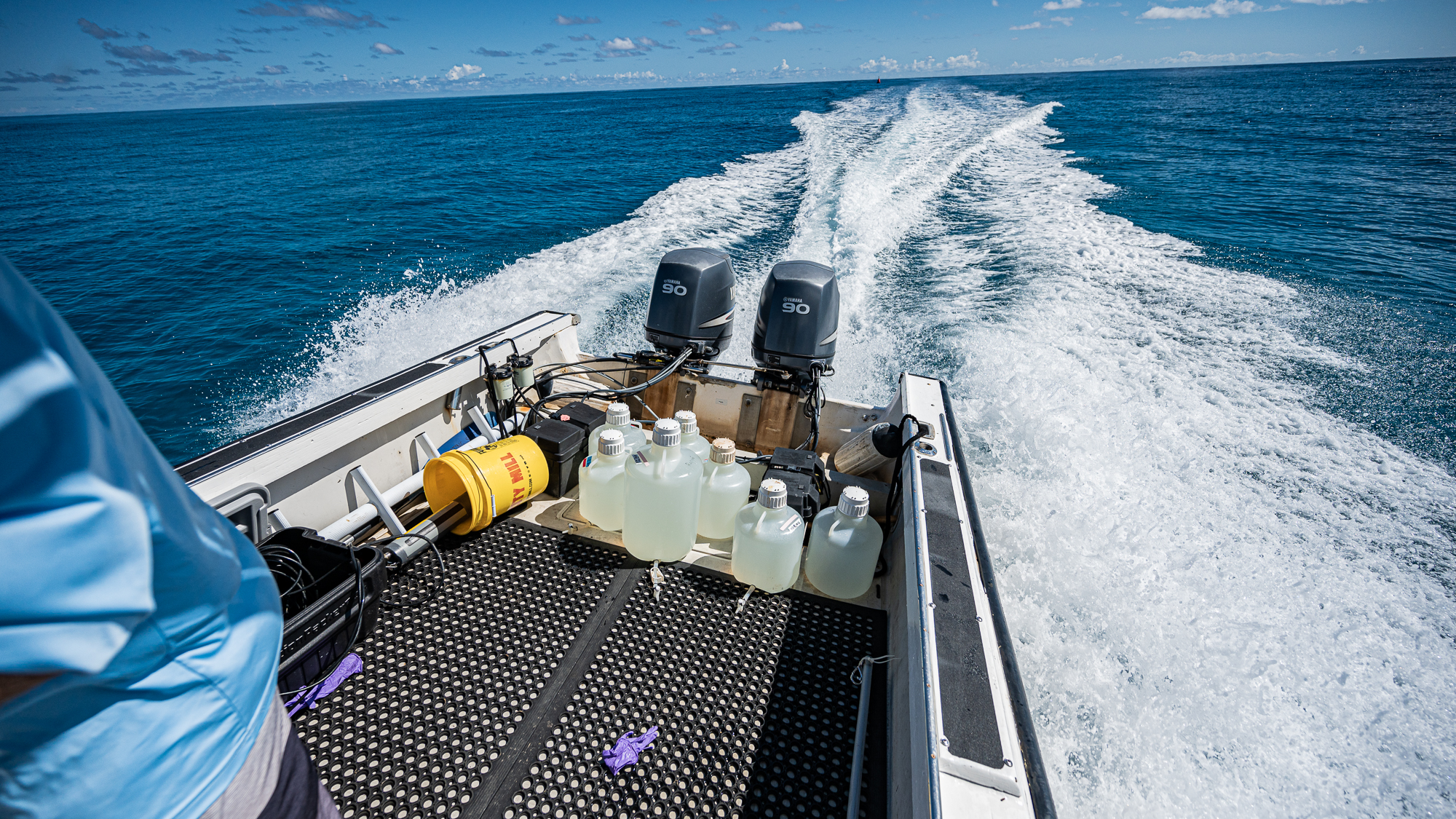
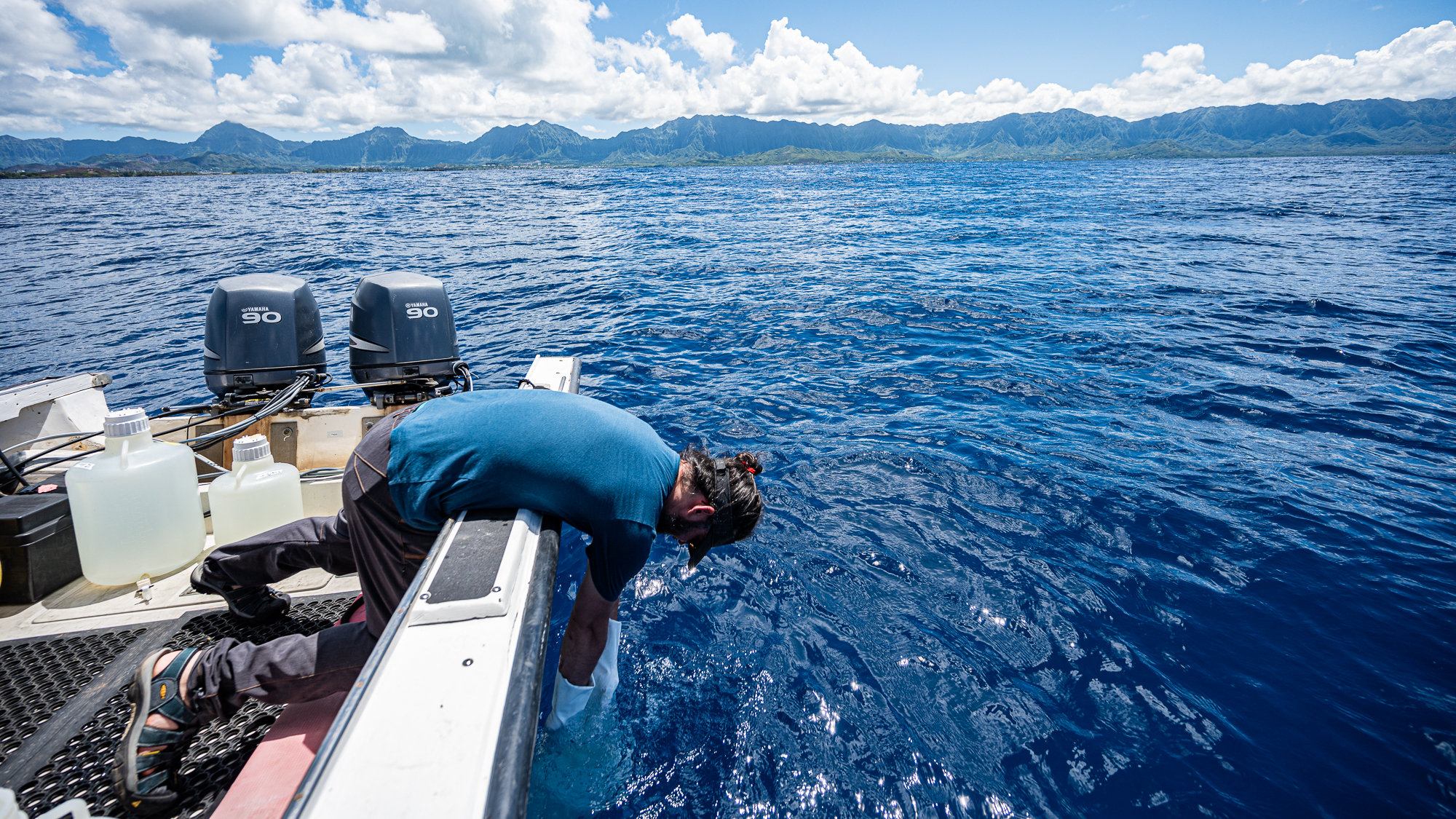
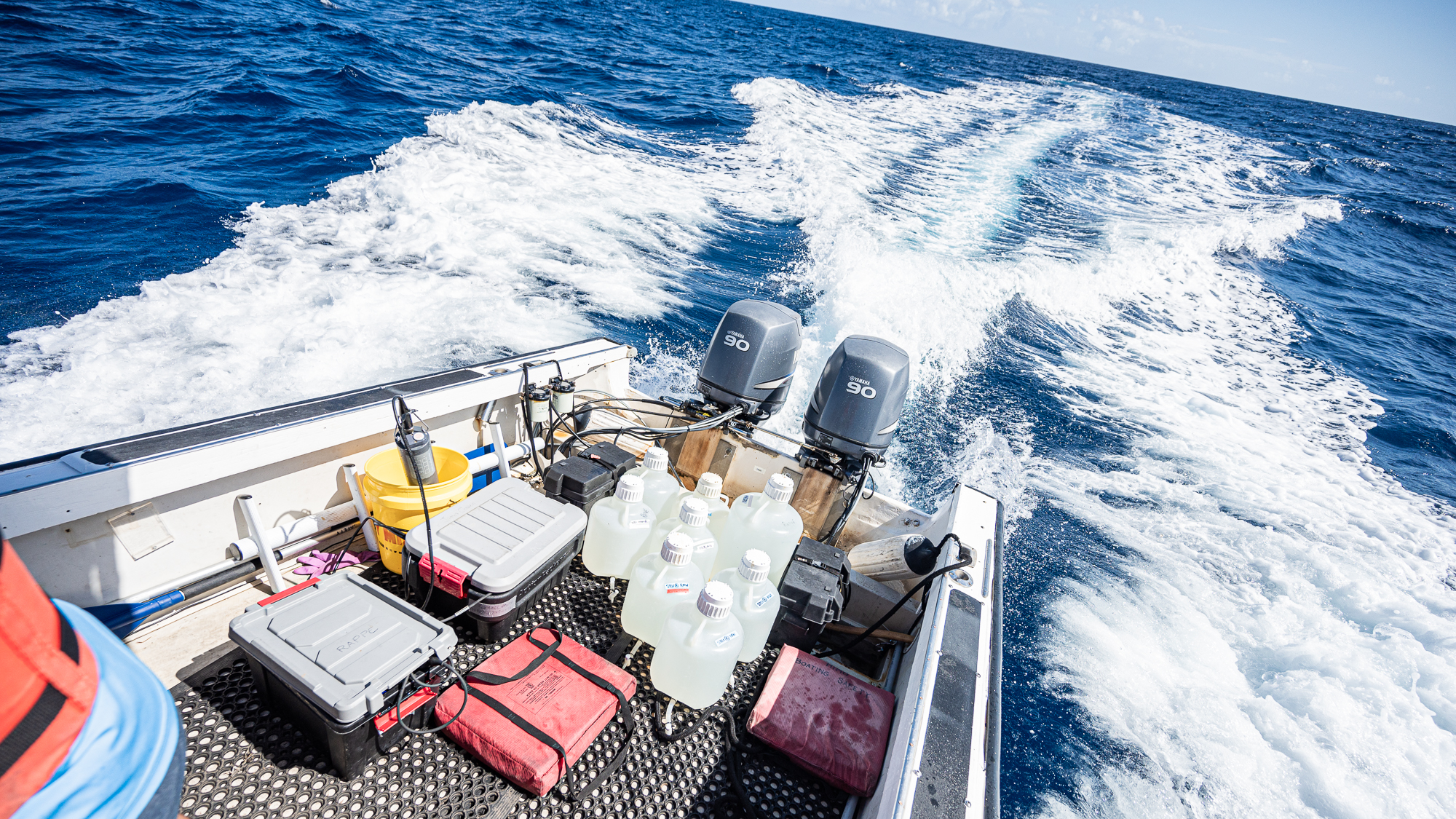
Mine was to sample the seawater by filling up 10L and 20L carboys at both stations.
It was difficult work: (1) filling one-fourth of each one of the 7 carboys to wash them three times exactly, (2) drowning them mercilessly, and (3) pulling them back in the boat when they had no air left to spare. The next day I was going to find myself covered in bruises at strategic locations of my chest where the boat met the meat.
But it was so worth it: this was the first time in my life I was physically contributing to the generation of samples that we were going to use to understand things later. With this realization, I gave my camera to Evan Barba, and asked him to take a photo of me so I could write to my mom to tell her “look, I finally made it!”.
And then I actually remembered this one exchange with a colleague that had really broken my heart. During a heated discussion a few years ago, an established colleague of mine, whom I still respect a great deal, had stopped me in the middle of my sentence to tell me that as a computer scientist I needed to consider leaving marine microbiology matters to marine microbiologists. Perhaps that was not a terribly bad idea. Yet here I was collecting surface waters from the Kāne’ohe Bay during a joint diel sampling effort with a great team of scientists, LITERALLY holding marine microbiology matters with my white dishwashing gloves. Of all the negligible wins I have accumulated throughout my life, I think this one was my true moment of arrival. One that no one was ever going to be able to take it away from me. Ever. My acid reflux disagreed, but I didn’t care.
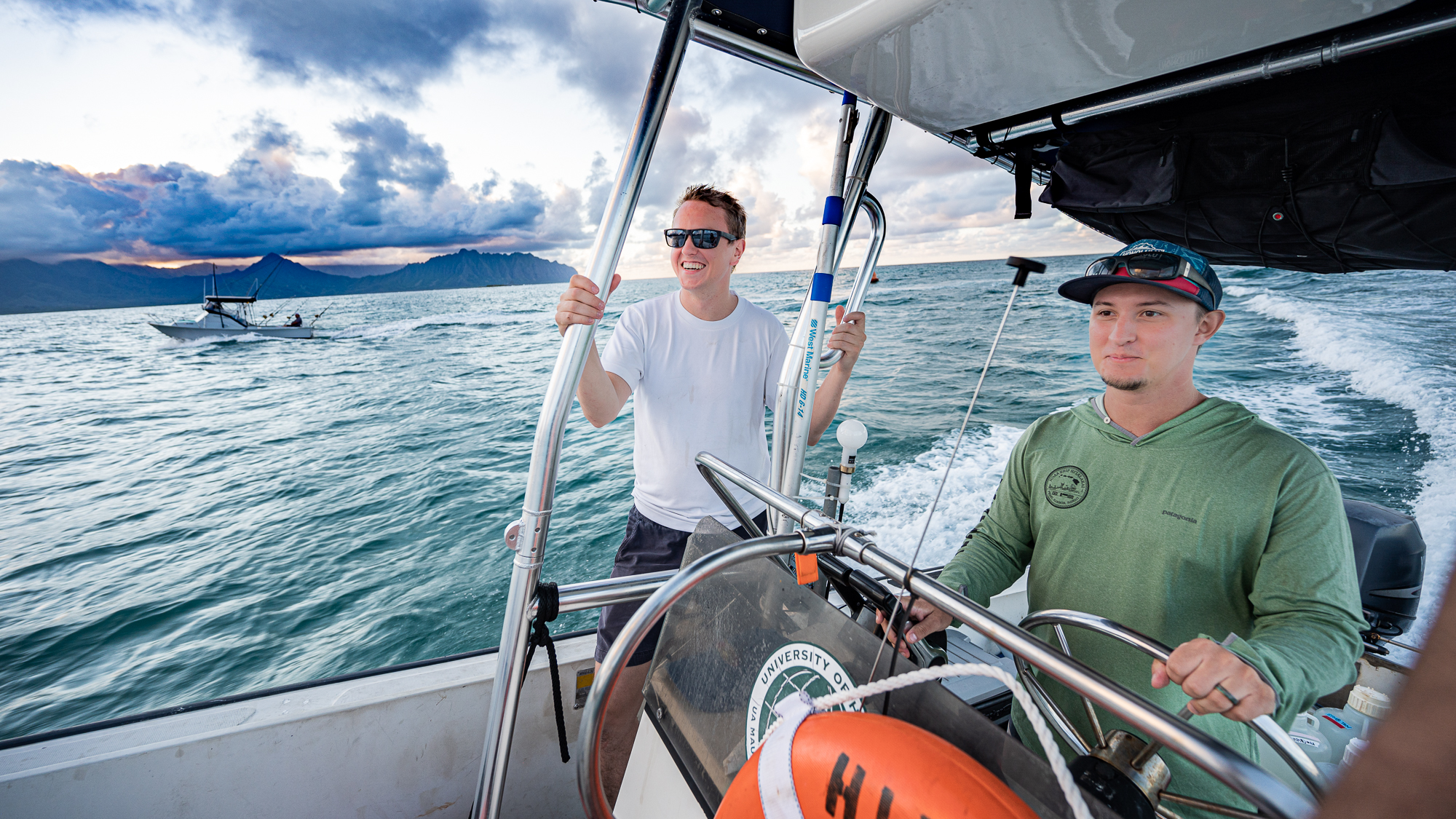
The sunset of the first day of sampling was approaching, and we were feeling pretty good about how smoothly everything had gone so far.
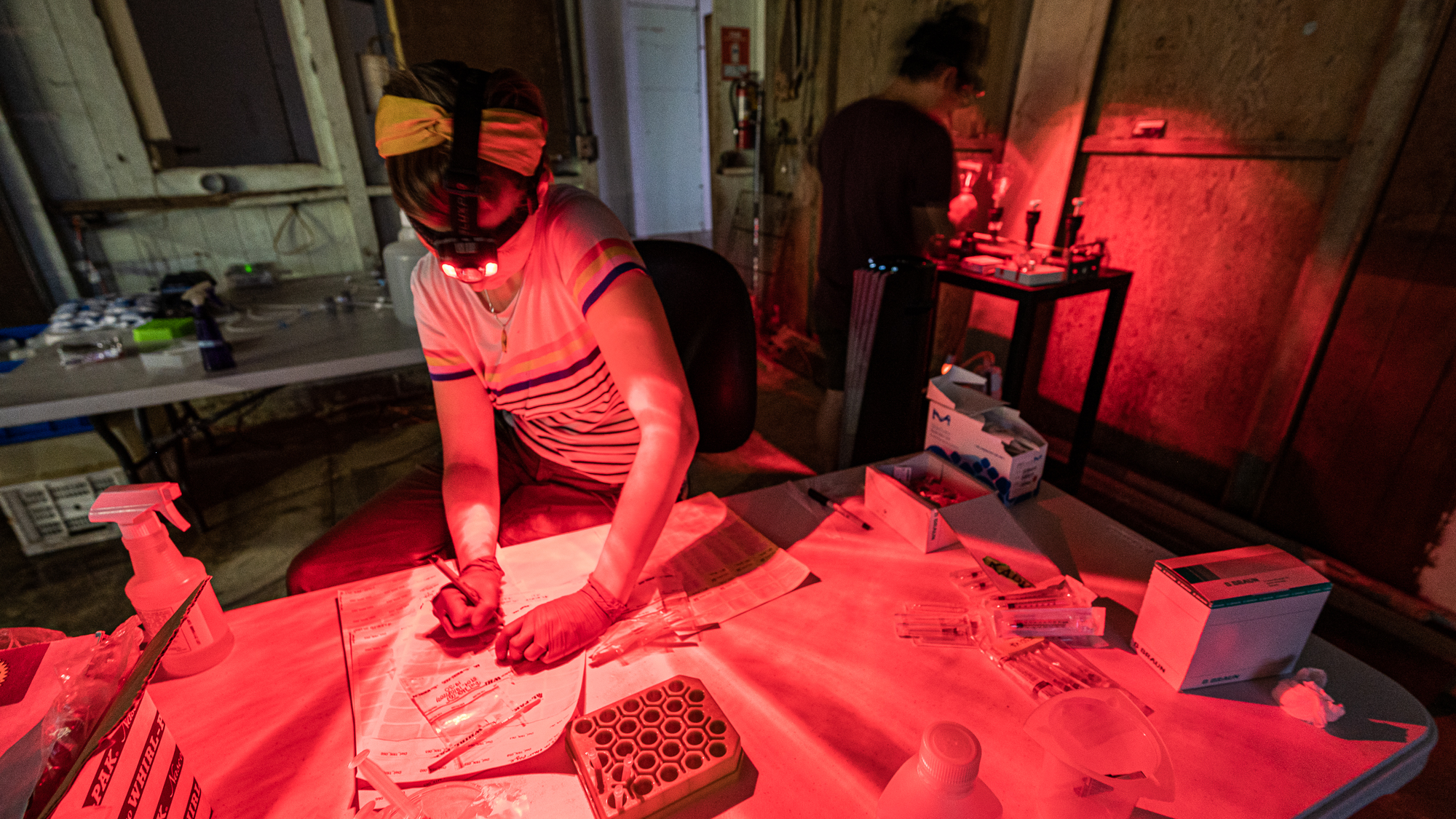
Meanwhile at the lab, Jessika, who was approaching the end of her 18-hours work day, was turning on her red headlamp along with others to not disturb the sleeping microbes.
Close to midnight I left the lab to get some sleep. Evan Kiefl, Clarisse, Mariana, Oscar, Adi, Mike, Mari, Sarah, and Kelle were taking over the shifts of the dark hours until the morning to drive the boat, collect samples, and process them.
August 19
When I arrived to the lab around 6am, I found Sarah, Mariana, and Mari working on yet another batch.
The sheet had many more lines now. Clearly things had gone well during the night shift.
Sarah did not waste any time to benefit from my astonishing wet lab skills and asked me to handle one of the most critical jobs in the lab: babysitting the spent bottles to measure the amount of filtered water per sample. I conducted extremely precise measurements of each bottle, without letting even a single drop of seawater go unaccounted for. I was very proud of myself.
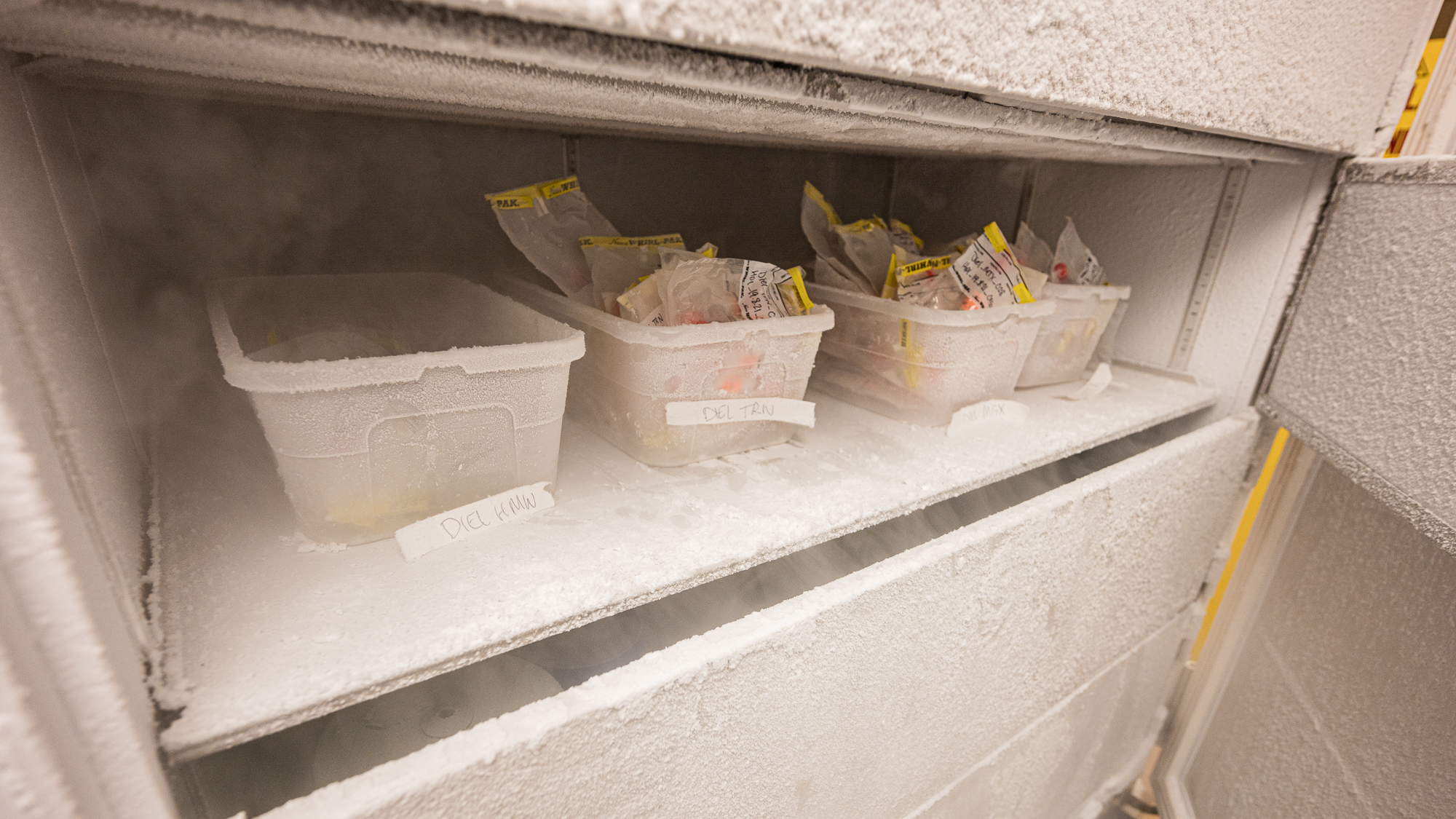
As filters were accumulating in the lab, they were quickly being carried to the -80C freezers in the building.
When I carried upstairs one of the filters Sarah was done with, I was quite excited to see the bins in the -80 that were slowly filling up with samples.
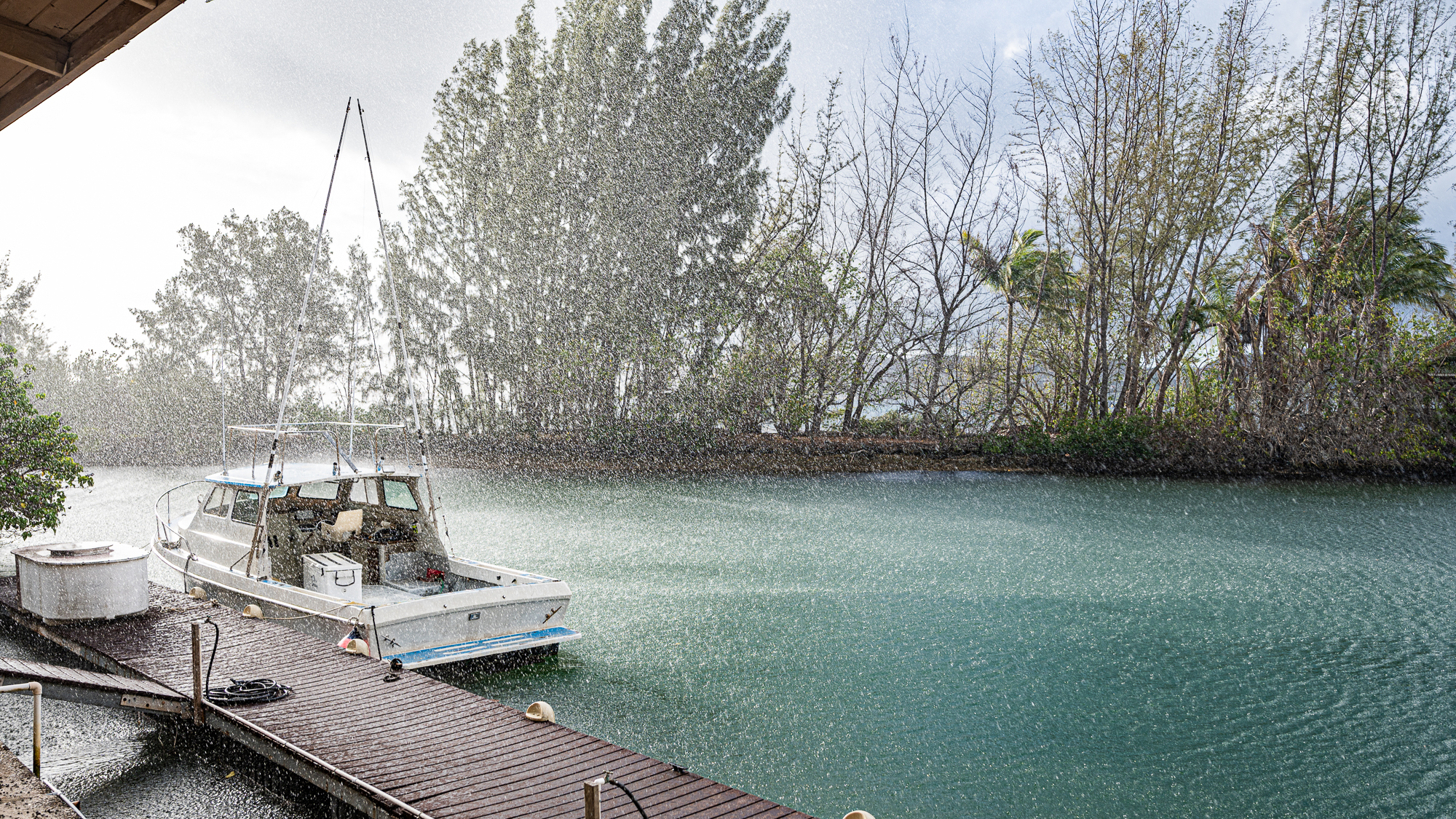
According to the schedule, my lab days were over. It was raining.
My shift at the boat for more sampling started with captain Kelle and Hannah.
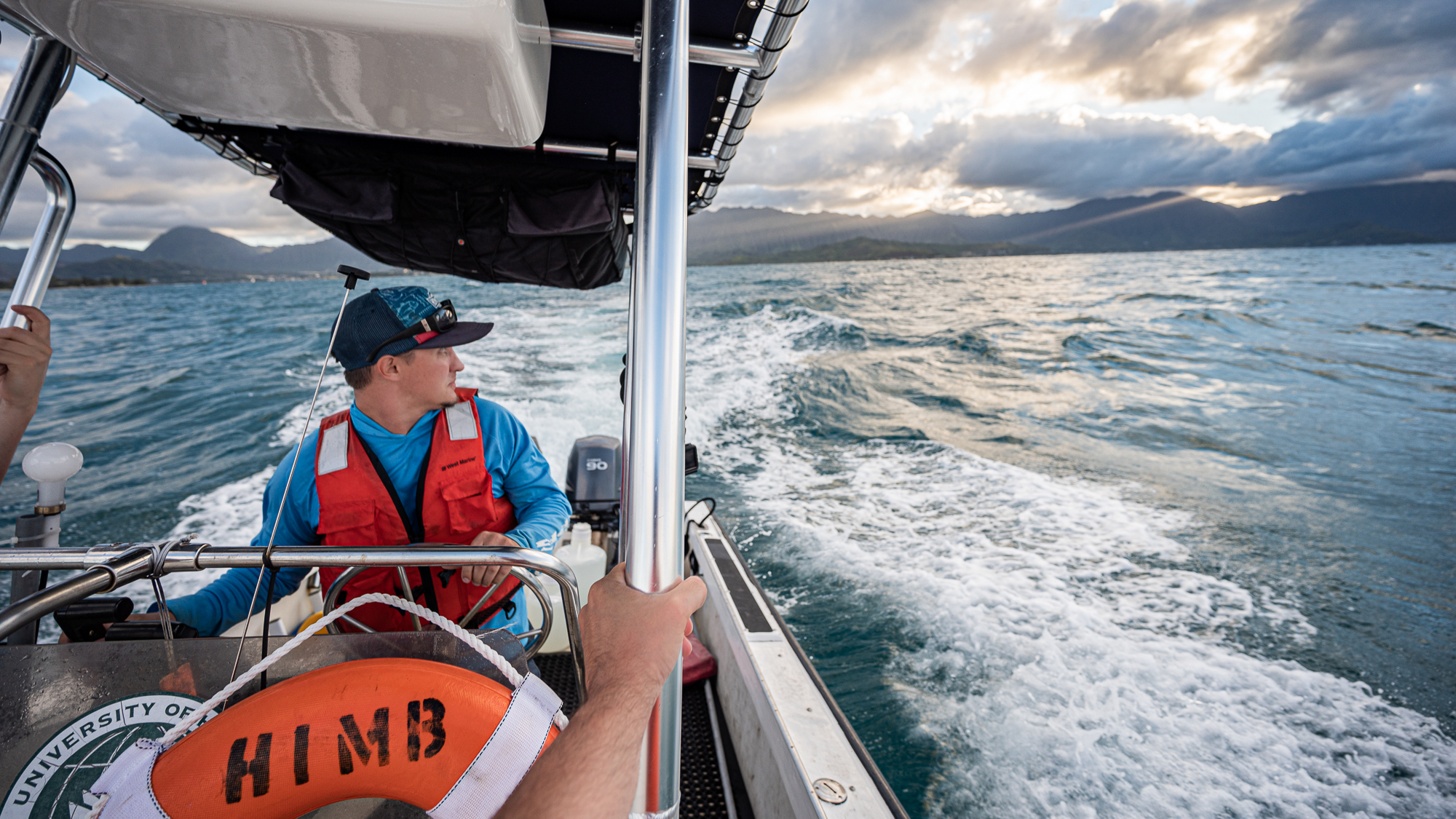
Today the weather was less forgiving. In addition to occasional showers, there was a lot of swell compared to the day before (as you can imagine someone was very happy and active given the situation).
But everything went well both at the lab and in the boat. My shift in the boat that started with Kelle was ending with Evan Barba.
After my last round, I bid my farewells to Adi and Jessika at the lab for one more night, and went my dorm room.
August 20
The final day of sampling started with a beautiful sunrise. Just like everyone else, I was extremely exhausted, but we were almost at the end of it. The only remaining rounds were to bring additional samples that could be used for testing purposes during library preparation.
Mike Rappé had been on the boat the entire night. While Evan Barba was giving him a ride to the parking lot so he can drive home for some sleep, Mike told us about his night and experience just like the day before. I felt like I could get used to this routine.
More than 48 hour after the beginning of this intense sampling experiment, we were finally bringing the very last water samples to the lab.
Every line in our sampling sheet was now populated with precious surface ocean samples sitting in Mike’s freezer. Except for minor hiccups, the sampling was a success.
We cleaned up our make-shift lab, and took the rest of the day off to rest.
Looking back, I am fully aware that none of this would have been possible without the ambitious early-career researchers, Evan Kiefl, Jessika, Sarah, Evan Barba, Kelle, who are photographed above, and all the others who volunteered their precious time and energy for this project. Surely there will be even more people will get involved with it as we go forward with the analysis of these data.
I am also thankful for the Hawai’i Institute of Marine Biology and its generous community that welcomed us.
And of course the Simons Foundation and the Alfred P. Sloan Foundation for providing us with the means to push our boundaries.
But we certainly could not come anywhere near where we are without Mike Rappé and his relentless support for this project since its inception.
Science is difficult. But its people make it worth it.
Next Steps
Well. On the one hand we have done a lot. Formalizing the idea, raising funds, planning, and completing an ambitious sampling effort to materialize the project took a significant amount of work. On the other hand, we are still at the beginning of this project. We still need to process our samples to prepare sequencing libraries, perform sequencing, analyze and interpret the data, and write our findings.
As I am writing these lines, we are back in Chicago. As samples come from the Rappé Lab’s freezers to us in dry shippers batch by batch, Jessika and Karen are working on extracting DNA and RNA molecules.
I will do my best to document the progress in this project to complete its entire journey from boats to bits. So in a way, this is to be continued.