The purpose of this blog post is to tell the story of how the scientific review process failed to capture the reproducibility issues in a human microbiome and autism study, and how we attempted to resolve this issue post-publication by investing a substantial amount of effort, just to fail ourselves at the end.
This post will likely speak to you if you are (1) an early-career researcher from any discipline who wishes to understand the role of scientists in the scientific review process to maximize reproducibility of science, (2) a senior scientist from any discipline who wishes to see a yet another example of how wrong can things go, or (3) a PhD student who really needs to write that paper but is also deeply convinced that procrastination is the only option at this time.
At the end of this post you may find yourself wondering “is there something I could do to fix this?”. Yes, there is. And in fact, in a Comment we published along with this blog post, we offer some 2 cents for you to consider carrying into your future activities as a peer reviewer.
This blog post is updated since its first publication. You can see precisely how it changed over time by clicking here.
We know. Reading long and gloomy things is not easy, or fun. So we also wrote this summary, which essentially is an easier-to-read TL;DR of this blog post, for those of you who wish to leave early but also want to understand what is this all about before that:
Once upon a time a study with fancy claims,
appeared in a journal of big names.
The journal required the data to be open and shared,
just like every other respectable journal that cared.
But of course it is one thing to 'require' something,
and another to enforce it as "well, we're just saying".
While the authors uploaded their data to a service,
they did it without the sample labels,
making it impossible for anyone to test their claims.
And neither the reviewers of the study nor its editor,
checked if the data were really there, or if it was just thin air.
And so it was Iva and Meren's burden to notice the neglect,
with the feeling that this was something they had to correct.
They wrote to the authors, who were unable to deliver.
They appealed to the editor, but not much was done there, either.
So they decided to turn their frustration and acrimony,
into something that could offer everyone a learning opportunity.
The motivation behind this blog post is to show you,
how something that requires a few minutes of a reviewer,
can turn into something that will remain forever beyond repair.
We often say "reproducible science is good and data must be open and FAIR",
but examples like this one actually demonstrate how much we really care.
Now this is out of the way, we hope you can leave in peace. For those who wish to stay, we have a lot to cover.
Introduction
As a research group that leverages ‘omics to study microbiomes everywhere, we routinely rely on publicly available data to contextualize our findings or search for new insights. Indeed, most of our published research uses publicly available data one way or another, and in return we spend a lot of energy to maximize the transparency of our work by making our data and bioinformatics workflows accessible. We find data and code availability to be a non-negotiable aspect of modern microbiology, as they not only support reusability of these critical resources that are often at least partially funded by public taxpayer money, but also keep scientists accountable for their published scientific claims. While most researchers and journal editors today would say they agree with this statement, in practice, many published studies fail to live up to this ideal of fully reproducible science through publicly-accessible datasets.
We repeatedly encountered significant challenges related to data accessibility and reusability in microbiology. These issues ranged from purely technical shortcomings, such as sample accessions and metadata buried in hard-to-process PDF files, to more troubling cases of mislabeled samples that required extensive data wrangling to diagnose and correct, or deliberately omitted links between sample names used in publications and sample names assigned to individual files in publicly available datasets. The latter form of obfuscation is really dangerous as it allows study authors to ‘check the box’ for making their data publicly available, while rendering these data largely unusable for secondary analysis efforts.
We have engaged in lengthy exchanges with authors to resolve these issues, yet in many cases we were still unable to access data or metadata that were essential to independently scrutinize published work and the claims within. In some cases authors have blatantly asked us to justify our intentions in detail regarding how we plan to use “their” data so they could decide whether or not to make them available to us, while in other cases they cited a lack of time or personnel to make datasets public. We were even pointed out that “it would not be ideal” for the authors to make datasets public yet, because they were still working on them for more publications. In all these cases the data or metadata in question had already been used to support published findings.
Once a study appears in the scientific literature, there is no valid excuse for keeping its underlying data behind closed doors. Studies that fail to enable other scientists to reproduce the work and test published claims without having to contact the authors should trigger mandatory correction tasks or conditional acceptance decisions from journals until the authors fully correct the data availability issues.
That is the only way for us to be able to tell society that science deserves to be funded by societies because it generates knowledge that can be scrutinized, reanalyzed, and corrected by anyone. We simply can’t have it both ways. But as you will see below, we actually do.
The study in question
The study we want to use as an example of occasional failures around scientific review and reproducibility is the one by Su and Wong et al., titled “Multikingdom and functional gut microbiota markers for autism spectrum disorder”, which appeared in Nature Microbiology in 2024.
It is a large study that focuses on the human microbiome and autism disorder, and a good example of “formulaic” work in science, as every single methodological component of it has been previously used in other high-profile microbiome studies. In that sense, it stands on pretty well-trodden ground. It also follows a familiar technical pipeline in the human microbiome field that has been applied to colorectal cancer, inflammatory bowel disease, liver disease, and more: find associations, build a classifier, validate, claim diagnostic potential. In this sense, the overall study does not substantially differ from a great number of publications taking this approach that frequently appear in the human microbiome literature. But it does have a large cohort, which is a true strength of this work.
The authors study 1,627 children from five cohorts and use metagenomic sequencing of stool samples to suggest that a panel of 31 microbial markers successfully distinguish children with autism spectrum disorder (ASD) from those who are neurotypical.
Their findings include a mechanistic narrative: the authors suggest that two biosynthesis pathways are depleted in ASD children (ubiquinol-7 and thiamine diphosphate), and they link the depletion of this metabolic potential in the microbiome to decreased antioxidant activity of microbiome and ultimately neural function of the host. The authors validate these insights across independent cohorts and report that the performance holds across ages, sexes, and geographies. Furthermore, the authors show that their panel does not flag children with ADHD or atopic dermatitis, and they suggest it is ASD-specific.
These are strong claims that require strong scrutiny. But unfortunately, an opportunity for strong scrutiny, one that may require independent analyses of the data used in the study to test these claims, is precisely what is missing here.
What is at stake
To keep the focus of our blog post on the structural problems this work demonstrates for reproducibility of science, we limited our scientific discussion to the framing of claims relative to what the study designs can support, rather than attempting to evaluate the scientific findings themselves. Readers who would like to learn more about the broader question of whether and how the human microbiome relates to autism may find the perspective by Kevin J. Mitchell, Darren L. Dahly, and Dorothy V. M. Bishop a good start: “Conceptual and methodological flaws undermine claims of a link between the gut microbiome and autism”.
There are some problems with the narrative of this work. For instance, the paper follows a causal logic chain often used in the human microbiome field that mixes causality with association. In this particular work that narrative goes like this: The gut-brain axis exists. Children with ASD have different microbiota compared to those that are neurotypical. Gut microbiome development is delayed in children with ASD. Transferring poop from ASD children to germ-free mice leads to ‘autistic-like behavior’. Transferring poop from healthy donors to ASD children improves their symptoms. THEREFORE, the microbiome contributes to ASD development, and microbiome markers can diagnose it. What does not come across immediately to a naive reader is that each step in this chain does enormous inferential work, and the leap from “associated with” to “contributes to” to “can diagnose” is not fully justified by the cross-sectional and observational study design.
This overreach that creeps into a “causality” narrative is not uncommon in the field, and not even a one-off with this group’s work. Here is a list of reviewer concerns for another study by the same group that was published in Nature Communications in 2026, which uses the same dataset as the Nature Microbiology paper (but because Nature Communications is more ambitious when it comes to transparency, you can read the entirety of reviewer comments and author responses in the Transparent Peer Review file at the end of this page and see how many times reviewers had to point out one key problem over, and over, and over again):
- “The statement that ‘cohabitation with ASD siblings worsens gut dysbiosis’ implies directionality and causation in a study that is entirely cross-sectional” (Reviewer #1, commenting on the initial submission).
- “The possibility of reverse causation, shared environmental exposures, or common genetic susceptibility is acknowledged only briefly in limitations and not reflected in the interpretations drawn in results and discussion” (Reviewer #1, commenting on the initial submission).
- “The conclusions about ‘shared pathogenic or beneficial strains’ across siblings lack adequate technical rigor and may conflate co-acquisition with actual transmission” (Reviewer #1, commenting on the initial submission).
- “There is a tendency to overstate causality, especially regarding environmental or familial influence” (Reviewer #1, commenting on the initial submission).
- “The study addresses the question about a potential pathogenic role of the gut microbiome in ASD, but fail to provide convincing evidence for a causal role of the microbiome” (Reviewer #2, commenting on the initial submission).
- “Results do not prove a causal role of altered gut microbial composition in ASD, and other than confirming the lateral transmission of gut microbial features” (Reviewer #2, commenting on the initial submission).
- “The results support some of the conclusions, except for the claimed clinical implications” (Reviewer #2, commenting on the initial submission).
- “Rephrase any statements implying influence, susceptibility, receptivity, or directionality to strictly associational language, as no causal inference can be made from the current study design” (Reviewer #1, commenting on the revised submission).
- “Strain sharing observed in a cross-sectional cohort cannot distinguish direct microbial transmission from co-acquisition or convergence and should therefore be framed as association or co-occurrence only” (Reviewer #1, commenting on the revised submission).
- “Labeling taxa as pathogens risks clinical overinterpretation, particularly when pathogenicity is context-dependent and not assessed in this study” (Reviewer #1, commenting on the revised submission).
- “Phrasing that implies intrinsic host susceptibility or biological receptivity is not supported without direct host-level functional measurements” (Reviewer #1, commenting on the revised submission).
- “Translational or interventional relevance cannot be inferred from cross-sectional observational data and must be framed as hypothesis-generating only” (Reviewer #1, commenting on the revised submission).
- “Host genetic contributions are discussed without corresponding genetic data and must be clearly identified as speculative” (Reviewer #1, commenting on the revised submission).
- “The manuscript should include an explicit and unambiguous statement clarifying that directionality cannot be inferred, as from a cross-sectional study, one cannot distinguish between direct microbial transmission, co-acquisition from shared environments, or microbiome convergence driven by unmeasured host factors, and no inference regarding directionality can be made” (Reviewer #1, rather valiantly commenting on the revised submission (even though their comments were not enough to catch the missing data, which we will get to later in this blog post)).
It is clear that the causal framing was so deeply baked into the work that even after a round of revision the manuscript still contained enough causal language to force Reviewer #1 to really spell it out for the editor and the authors at the end (which finally appears as a Discussion paragraph in the published work). Whether deliberate or habitual, the end result is the same: a specific narrative that is more wishful and compelling than what the actual data and analyses can support. Nature Communications was lucky to have a fair reveiewer in this case who recognized the good bits of the work, but also had the expertise, the rigor, and the time and energy to actually push back on the poor bits of it over and over. But this is not always the case, and in fact, in journals with less rigorous review, the same group turned similar claims into published claims. After all, in some ways this is a numbers game: if you try enough times, you will find a journal where Reviewer #1 is not in the room.
For instance, in their 2026 study that was published in the journal Gut (by BMJ), the authors openly claimed that gut bacterial genomic structural variants are linked to gut metabolic dysregulation and dysbiosis observed in children with ASD. Given that structural variation in genomes (from any domain of life, not just bacteria) do not imply any functional consequences, associating the presence or absence of those variants to host health does not link them to disease in any way. The authors themselves admit the limitations of their work in the manuscript, but such statements of limitations are buried deep, and remain only accessible to those who have the skills (and time) to read the entirety of the publication. In contrast, the clear statement of linkage appears in the title page of the paper, right under the section “How This Study Might Affect Research Practice or Policy”; the part of the work that is most accessible to all members of society by design.
Another 2026 study by the same group published in the journal International Journal of Molecular Sciences (by MDPI), went much further, and managed to shove this title into the science literature: “Gut Microbiome Mediates the Causal Link Between Autism Spectrum Disorder and Dietary Preferences”. Besides the absurdity of this work that generalizes from middle-aged British adults to juvenile ASD populations elsewhere, the study objectively does not establish causation of any kind. Once again, the authors included statements in more technical parts of the paper that describe the limitations of what they did, but “causal link” is in the title, which is the part that gets indexed, that is cited, and that is read by families, doctors, investors, and/or policy makers, who may or may not see a difference between an MDPI journal and Nature Microbiology.
The strategy of asserting causality in the framing, retreating to association in the fine print does appear even at the highest levels of the human microbiome field. But admittedly it is rarely this blatant, which is somewhat refreshing in a tragic fashion, since it makes it easier to point out the dangers of this rhetoric.
But why?, you may ask, why push towards causality so determinedly if the current evidence doesn’t support it? To answer that, we invite you to read the vision statement of InnoHK Microbiota I-Center (MagIC), a microbiome research center and the employer of 12 of the 16 authors of the Nature Microbiology study. See if you can catch the “we use impressive technology” -> “we can diagnose things” -> “we can develop personalized microbiome therapeutics” arc:
That is where the causality language becomes critical. Because without it, this arc would look less like the Arc de Triomphe in Paris and more like an pigeon nest in New York. As a result, the kinds of products the authors are promoting may fail get the kind of support they need to justify their existence:
A critical perspective here may rightfully propose that these are all subjective concerns that require additional analyses of data to substantiate. Yes, indeed. And that is precisely the point of this blog post. The Nature Microbiology study prevents such a discourse as there is no practical way for an independent researcher to test the core claims using the data released with the study. Because the Nature Microbiology study, the Nature Communications study, and others by the same group that explore associations between microbiome and autism all end with a Data Availability statement that reads along the lines of “participant metadata cannot be made publicly available […] to protect participant privacy”.
We are all for privacy. Details that can identify patients should not be publicly available under any circumstance. But anonymized sample metadata (i.e., ‘which sample is from the neurotypical group and which is from the ASD group’ in this case) is not that. These group labels were not available when the Nature Microbiology study was reviewed, and they are still not available to those who may want to review it further to explore the authors’ claims. This blurs the ‘reproduciblity’ line that we often point to in order to give science more credibility in the case of this study.
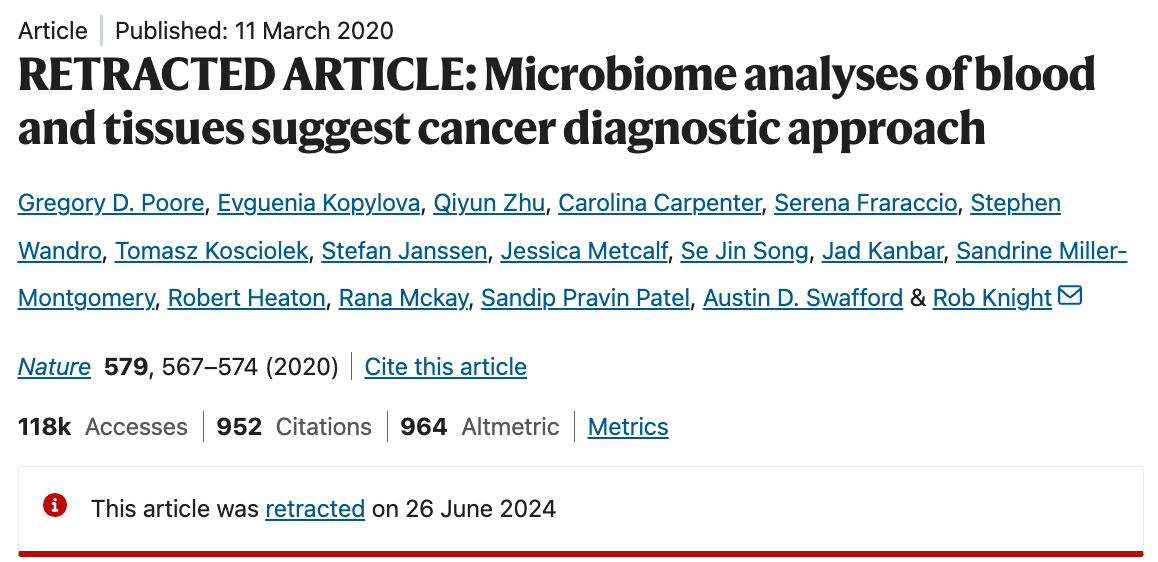
If you are still not sure why this is an important point, consider this: the ‘human microbiome <> autism diagnosis’ narrative is eerily similar to the now-retracted paper on ‘human microbiome <> cancer diagnosis’ narrative:
The retraction came as a result of a rigorous post-publication analysis conducted by Gihawi et al., who would not have been able to do such an analysis if the original data or cohort labels were not available, which is the case with the Nature Microbiology study.
What does this mean for families who are desperately searching for solutions to help their loved ones with ASD?
By publishing this work, one of the most prestigious journals of our field is effectively putting a stamp on a diagnostic narrative that dabbles in therapeutic potential of the human microbiome in ASD. But who can the families turn to to understand the implications or potential shortcomings of this work? One of the most powerful arguments that justifies science spending in the public mind is its openness to scrutiny. We tell them, you can trust science because it is easier to scrutinize than most other things you are told.
But who really gets to scrutinize science? An average human gut metagenome contains hundreds of millions of sequences that can be analyzed in tens of different ways that require not only a significant amount of computational resources but also years of expertise. So, who can really argue with the claims of a study that analyzes hundreds of metagenomes to support its narrative, if not other scientists with more or different expertise? Given the technical complexities of scientific data, the public is effectively forced to put their trust in the hands of other scientists. And scientists often prove worthy of that trust. For instance, it worked perfectly well with the cancer microbiome paper: the underlying data were available for scrutiny, and that benefited everyone, including the authors of the original work who made sure that their study could be scrutinized. But when data are not available to other scientists, claims that can turn into financial gains through ambitious companies or that can change power dynamics through new policies remain unchecked, eroding the very reason why science exists and what it represents.
Timeline of events
Welcome. This is the story of how we tried to be good scientists and scrutinize some strong narratives, and how we failed.
Jul 10, 2024
The paper by Su and Wong et al. appears online.
Jul 18, 2024
We (Iva and Meren) discover its existence, and start studying it with great interest.
While downloading the metagenomes, we realize that even though there are only 1,017 metagenomes online, the ‘discovery cohort’ of the paper includes 1,083 children, the ‘independent hospital cohort’ includes 172, the ‘community cohort’ includes 176, and so on. Numbers don’t match, and it is not a very good start. But genuine mistakes are a part of science. So that’s that.
Jul 24, 2024
We go to the GitHub repository that is linked in the paper at qsu123 / ASD_multi-kingdom_diagnosis to ask about this, and discover that there is already a question about data availability by MengQingren, which reads,
Dear Doctor,
Could you provide the sample grouping information, such as whether S0001 corresponds to ASD or ND? I am looking forward to your reply.
This is not good news either. Without group information, none of the comparisons they make can be accurately tested.
Iva writes under the same GitHub issue, seconding this request:
The publicly available sequences at PRJNA943687 are an excellent resource for the community and could be useful for investigating several hypotheses if the correspondence between samples and their groups are known. We would appreciate the sharing of this essential metadata.
Aug 4, 2024
10 days later, we still have no reponse from the authors. Now cssmillie joins the conversation:
We are also interested in obtaining the sample metadata for PRJNA943687.
@ivagljiva @mengqingren @meren were you able to obtain it?
We report our failure.
Aug 5, 2024
GitHub repository owners get emails when an issue is reported or when there is a new comment on existing issues. Nevertheless, we decide to send an email to multiple authors of the study:
Dear Drs. Su, Wong, Chan, and Ng,
We recently read your study in Nature Microbiology, Multikingdom and functional gut microbiota markers for autism spectrum disorder. Your finding that functional and metabolic factors differentiate the gut microbiomes of neurotypical children and those with ASD is very compelling, and it was also nice to see a machine learning study with careful model validation. Congratulations on the great work :)
The public data you have made available appears to be missing sample group information (ASD vs neurotypical) for the metagenomes in your study. There is some sample group information in your GitHub repository, however, the sample names therein are different from the sample names in the SRA metadata for PRJNA943687, which makes it impossible to understand how the public data corresponds to the data used in the study.
Would you please consider making them available? We appreciate your consideration and are looking forward to your response.
Best wishes, Iva & Meren
Another 24 hours go by without any response.
Aug 6, 2024
Meren decides that it is time to reach out to the Journal and writes to the Chief Editor of Nature Microbiology to explain that,
-
The study lacks sample / group identifiers.
-
The public data by the authors does not include all samples used in the study.
The email ends with the following statement that summarizes the severity of the issue:
In its current form, the study is by no means reproducible. It is a pity that these glaring issues were missed during the review process, but the problem is not impossible to address. That said, with our inability to get a response from the authors, we are not sure how to proceed. The most critical need here is a proper metadata file that explains sample groups for publicly available data. The authors should not make their data available just to check a box to pretend that they satisfy the journal policies while leaving the data in a state that makes it impossible to reproduce their findings independently. We are wondering if you would consider this as something the Journal could help with.
Aug 7, 2024
The editor responds kindly and quickly and agrees that this is an important issue to address. They mention that they will immediately contact the authors and ask them to address these points as soon as possible.
Aug 7, 2024
Just hours after the editor’s email, the authors respond to everyone on GitHub:
@meren @cssmillie @ivagljiva @mengqingren Hi all, I am so sorry for I didn’t notice the comments here. We are now updating the PRJNA943687 and the metadata will be available soon. I will let you know when it is ready. Sorry again for any inconvenience caused by this.
This is great news, but as you can tell by looking at the scroll bar on your screen this is not the “and then they lived happily ever after” moment as there is more to go through in this blog post.
Aug 8, 2024
The editor writes back and confirms that data for some samples were indeed not included in the original upload, but this has now been fixed along with the metadata.
We are very happy to hear that the data and metadata issues were now resolved. But nothing really happens in reality, and metadata remains unavailable. We decide to wait a bit, and not burden the authors too much.
Sep 26, 2024
After six weeks of waiting, Iva finally writes on GitHub:
Hello @qsu123, just checking in to see if the data and metadata are available yet? :)
Sep 27, 2024
We get a response:
Yes. All the metagenome were released in the PRJNA943687 last month. Please contact the Corresponding author, Prof. Siew Ng (siewchienng@cuhk.edu.hk) for group information.
There should not be any need to contact anyone for the group information that was promised to be public. But we write the following email:
Dear Prof. Siew Ng,
We are interested in using your public dataset of gut metagenomes under BioProject PRJNA943687. Could you please share the sample group information (ASD vs neurotypical) for these samples?
Thank you so much for your help!
Oct 2, 2024
Instead of Siew Ng (the corresponding author), we get a response from Qi Su (the first author who told us to write to the corresponding author):
We are happy to provide the group information, but we need more details as below.
According to the current data sharing policy, requests for data sharing can be submitted with a written proposal. The proposal should detail the intended use of the data with reasonable methodology and sample size estimation. These requests are reviewed based on scientific merit and ethical considerations, including patient consent, to avoid any potential misinterpretation or commercial misuse. Additionally, please ask your supervisor or mentor with an academic position of assistant professor or above to submit the request. Since we are working with numerous data sharing inquiries from various individuals, it takes some time before the data is released, but we will make sure to do our best to complete it quickly.
Thank you for your understanding. Please let me know if you have any questions.
Sigh.
There are many ways to make data available without infringing patient privacy or violating any ethical considerations. Thousands of studies do that by anonymizing samples. This is not rocket science. Just your normal science.
But regardless of the reason, this is a failure of study design that should not turn into a failure of science by letting papers without data appear in Nature Microbiology, Nature Communications, or any other respectable journal.
One of the details that makes this ‘application for labels’ requirement questionable is that it asks requests to be submitted by someone with an “academic position of assistant professor or above”, suggesting that PhD students, postdocs, or independent researchers may not be able to independently scrutinize published findings on equal terms.
Of course this statement does not mean that the data are going to be unavailable to everyone. It just means that there is enough red tape around it to keep it unavailable to some. And this kind of structure for ‘access control’ can affect independent scrutiny and its severity. Not a good light to be under.
Oct 2, 2024
Upon this response, Meren reaches out to the editor again (carried in here with some redactions):
The authors seemed to be enthusiastic about sharing the data along with labels as required for reproducibility, but this was their response:
[copy-pasta of the author requirements]
I wanted to ask your input again regarding how Iva and I should proceed.
On the one hand, after months of struggle I am ready to write a ‘proposal’ just to revisit the claims these authors published in Nature Microbiology.
On the other hand, anyone can claim ‘patient consent’ or ‘potential misinterpretation’ to not share anonymized labels if they are not a fan of the idea that others may scrutinize their findings. Currently this paper represents everything that is wrong with data availability and transparency in science.
The editor quickly confirms that they will address this with the authors.
Nov 27, 2024
After about 8 weeks, the editor writes back, but they don’t have good news.
They let us know that the authors “cannot share metadata associated with this cohort due to the nature of the patient consent forms”. As a solution, they are working on a correction for the paper with an updated Data Availability statement, which will “include information on how to request” the missing information.
A few weeks later, precisely on Dec 04, 2024, the article’s data availability statement indeed gets an official update. In its final form it no longer claims that all data are available, but carries itself with a bit more flair:
Where does this leave us? Or the reviewers of this study? Or the public?
Here is a summary of what happened so far to ponder about these questions:
-
The authors claim that they have generated a consent form that is incompatible with modern scientific practices in medicine, such as ensuring that anonymized data can be made available to others to independently test the validity of scientific claims.
-
They nevertheless submitted their work to a journal that promises transparency and reproducibility of the science it publishes by requiring a means for readers to independently test the validity of scientific claims. But the study was initially reviewed and published with a Data Availability statement that did not disclose the practical limits on access to the key labels, which was incompatible with the journal policy (and that is precisely why they had to revise it with an official update to the already-published paper),
-
With the upated Data Availabilty statement, the journal went from ‘the data is publicly available’ to ‘send your requests to the corresponding author’ so that the data requests can be evaluated in a case-by-case basis.
Basically, the paper was reviewed in the journal with the assumption that data was available, ended up getting published with unfalsifiable statements, and it got to stay in said journal in its current form, despite the continued absence of essential data labels.
At its core, ‘open data’ principles are here to ensure that data generators have no means to prevent others from finding answers in their data that may contradict their interpretations.
It is worth thinking about whether the reviewers of this study would still have agreed to put their stamp of approval on this work if they knew this is what it takes to simply scrutinize its narrative? It is important to note that these statements did not appear in the official data availability section of the paper when the work was first reviewed. While the work was under review, and until this point, 21 weeks after its publication, the official Data Availability statement simply claimed that all data were publicly available.
At this stage we had this tingling feeling that perhaps we were not going to get these labels. But it was too late to let this go, so we did the only thing we could do: ask.
Dec 2, 2024
Meren writes a formal request as instructed, with the letter head and everything:
The letter is short by design. Its entire purpose is to simply say “we have no commercial interest & we are here to scrutinize your findings”. Here is the meat of it:
The intended use of the data and labels is to reproduce your findings and confirm the validity of markers you have identified in the aforementioned manuscript using alternative bioinformatics approaches and computational strategies. We have no commercial interests, and the scope of our investigation will be limited to the scope of yours.
This is indeed not the most comfortable thing to read for any author. Especially when it comes from a group that is pretty serious about secondary analyses of public data with decades of expertise in using computational strategies and metagenomics to study the human gut microbiome. But science is not a place of comfort. It is an exercise in learning over and over again that we can be wrong, and we need each other to check on one another. So in an ideal world, there should be no way to refuse such a request for data that appears in a currently published paper in one of the most well-respected journals of a field, especially after promising in said paper that labels will be made available upon request.
But of course, the language the authors used in their promise leaves enough room to refuse requests for vague reasons. The most obvious one is the requirement for ‘scientific merit’. One of the first things we learn as scientists is that the same grant application can be found to be completely devoid of any scientific merit by one review panel, yet it can be found to have the highest of all scientific merits by the next one. Scientific merit is a requirement here thanks to its extremely subjective nature. It enables one to not honor a given request, as anyone can accept or refuse the same request using ‘scientific merit’ as a justification.
Dec 19, 2024
About two weeks later, Qi Su kindly forwarded the following response on behalf of the ‘committee’ of unnamed individuals:
There is insufficient information in this application for further evaluation, such as the study objectives, research hypotheses, detailed methods, preliminary data, sample size estimation, and proposed conclusions. A detailed protocol is required to meet the needs of ethical and scienticial assessment. In addition, the applicant should clearly state whether there is a possible commercial application of their findings.
Here is a point-by-point response to these claims in the spirit of imaginary arguments we always win:
-
The desire to scrutinize one’s published claims is more than enough justification.
-
Not only is requiring access to “study objectives, research hypotheses, detailed methods, preliminary data, sample size estimation, and proposed conclusions” not appropriate, this is not in line with the official Data Availability statement, which does not mention that one would need to disclose all of these potentially sensitive details for the labels of a published study to be made available. But is this even surprising at this point? First we were told that all data was publicly available. We realized it wasn’t. Then we were told that the issues with data and labels were now resolved. We realized they weren’t. Then we were told that the authors would make the labels available if we send an application. Now we can see that they wouldn’t. Layers and layers and layers of this. Of course no one can keep up. Editors, scientists, clinicians, we all have other things to do than going after data availability issues for eternity, which makes these layers of bureaucratic tape a very effective strategy to avoid scrutiny.
-
And here is a fun one: the committee says “the applicant shoudl clearly state whether there is a possible commercial application of their findings”. Well, clearly there is ‘possible commercial application’ of every finding, regardless of whether the findings are falsifiable or not. If this is not clear to the committee, they can perhaps reach out to the authors who conducted this very study and explored ways to turn their unfalsifiable findings into a commercial application:
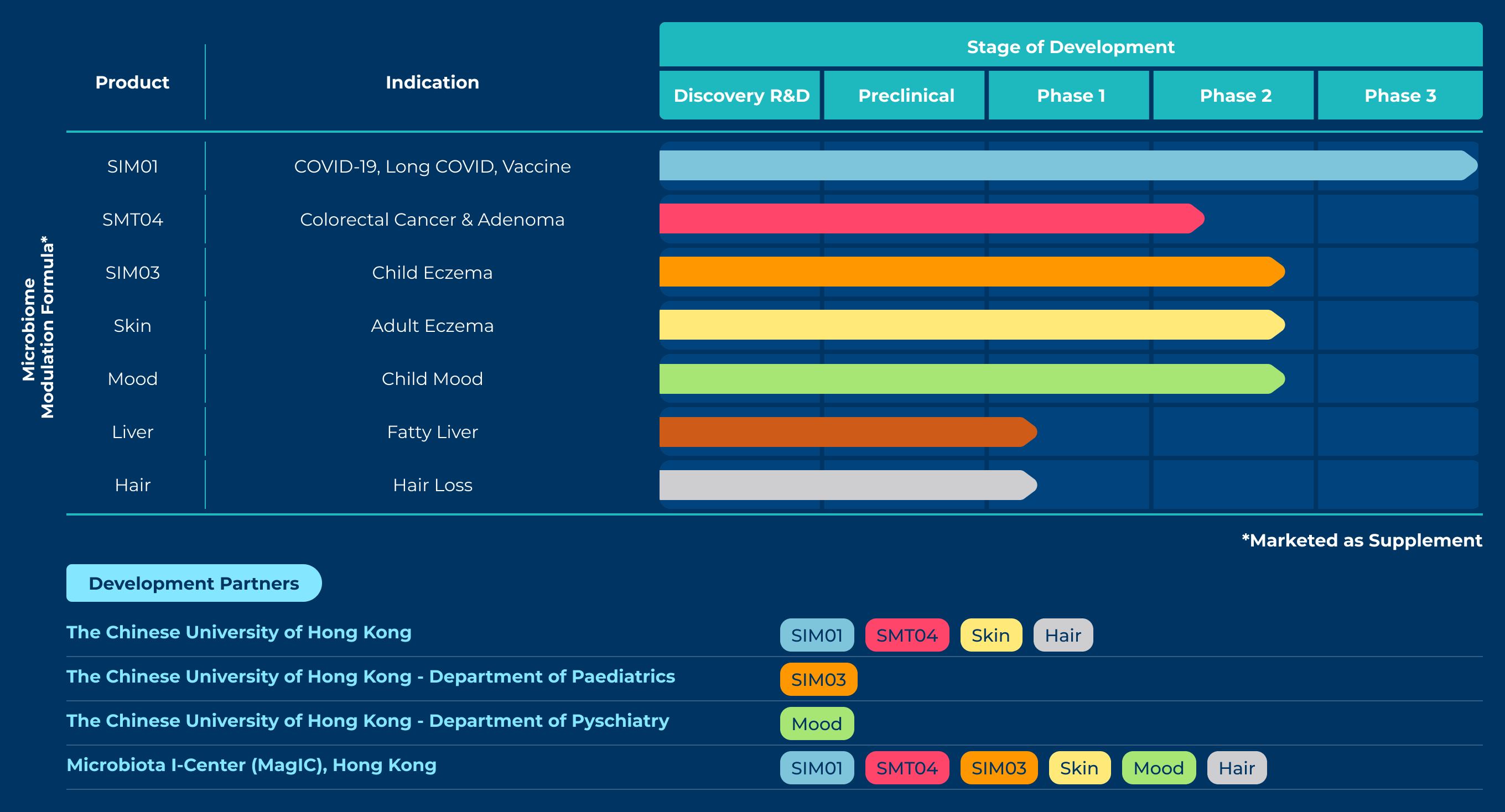
The GenieBiome Ltd listed here is a therapeutics company co-founded by Siew C. Ng (S.C.N.), the senior author of the Nature Microbiology study and the person you need to send your ‘application’ to for access to the data labels. The company says on its web page that it is “transforming a human first, data-driven microbiome discovery platform to drug development for multiple diseases”. Then lists all these products they are currently developing, with an asterisk at the corner that says “marketed as supplement”:
If the journey of this narrative that goes from ‘transforming’, ‘discovery’, ‘drug’, ‘disease’, all the way down to ‘marketed as supplement’ seems familiar, that is because it is not much different than the previously-mentioned strategy of asserting causality in the framing and retreating to association in the fine print. But get this: one of their products that is in its Phase 2 development is an “oral live microencapsulated bacteria for child mood”, which, according to the web page, reduces (at various levels of success) ‘anxiety symptoms’, ‘sensory hypersensitivity symptoms’, and ‘functional abdominal pain disorder’ in children with autism.
All these exciting statements cite a single source. That source is a press release that goes through the claims that are now published in the unfalsifiable Nature Microbiology paper. So the circle is complete.
If you are a parent, wouldn’t you give a chance to these products when they are promoted by scientists and approved by other scientists? Or, would it change a parent’s mind if they knew that other scientists could not test any of these claims because the people preparing to sell them these products for their children to literally consume are also those preventing others from accessing the labels of a scientific study that ties it all together?
Big sigh.
The only silver lining of this outcome was that it concluded our suffering. With this final response, we were done, and the scoreboard read,
Su and Wong et al. .......: 1
Iva and Meren ............: 0
But we had one last defeat to register under our names before the end of all this.
Jun 6, 2025
With our recognition that all of this could have been avoided if the reviewers of the Nature Microbiology study had paid just a bit more attention to the contents of the study they were reviewing, we thought that a gentle reminder for reviewers with an easy-to-follow checklist might be a good idea to submit to Nature Microbiology.
So we submitted “A data-conscious checklist for the reviewers of ‘omics studies who champion open science” on this date for the consideration of the editorial board to appear in Nature Microbiology as a Comment.
Aug 18, 2025
It only took about 10 weeks for us to get the rejection notice from Nature Microbiology, which said the editorial team felt that the comment would find a more appropriate outlet in another journal (i.e., not the one that published a paper after its own reviewers missed the glaring issues with its data availability and forced us to embark on an unwinnable quest, but another one). We tried Nature Communications (the other journal where the reivewers let another autism study by this group to appear in the literature without data to reproduce their claims), and we were rejected from that one, too.
Sometimes such is life in science.
Apr 15, 2026
Well. Nearly 8 months went by since then, and here we are writing this blog post, so this tale is not completely lost.
It takes time to get over these things. Especially for those who consider the implications of these failures, and their meaning for how we scientists see ourselves and how we are seen from the outside.
Our Comment is now on metaRxiv. Please read it, take a look at our reviewer checklist, and consider sharing it with your friends, colleagues, trainees, or editors of your favorite journals:
Final words
We started this journey because we wanted to test a hypothesis against someone else’s data: a most ordinary thing in science.
Instead, we spent countless hours writing emails, filing formal requests, and appealing to editors, only to learn at the end that the simplest question, “which samples are ASD and which are neurotypical?”, would not be answered, even though that very question has been the basis of a research claim that remains published.
The reviewers who could have caught this did not catch it. The journal that could have enforced its own policies did not fully do so. And the authors who could have resolved this at any point did not make the labels available in a form that enabled independent scrutiny.
We don’t know if their findings are right or wrong .. and that is precisely the point. Nobody does, and nobody can find out. If that is acceptable to us, then we should stop pretending that we care about open data and reproducibility. If it is not acceptable, then something has to change. And you, the mighty reviewer, can be a part of that change.
Data transparency is not always enough to tease apart good science from bad science. But it is all we have. So the next time you open a manuscript to review, ask yourself before anything else: “is it possible for anyone to falsify the claims in this study using the same data that was used to justify them?”.