Table of Contents
As you know, hierarchical clustering in anvi’o (that is necessary to represent a merged dataset as a nice looking tree) requires the maximum number of splits to be around 20,000.
If you have more, one way to do it is to profile your dataset with an --min-contig-length value that eliminates enough short contigs from your analysis so you can merge everything without any issues.
But what if you do not want to use a large --min-contig-length value and lose a lot of contigs that may increase your completion score?
Here is an example project with more 250,000 splits:
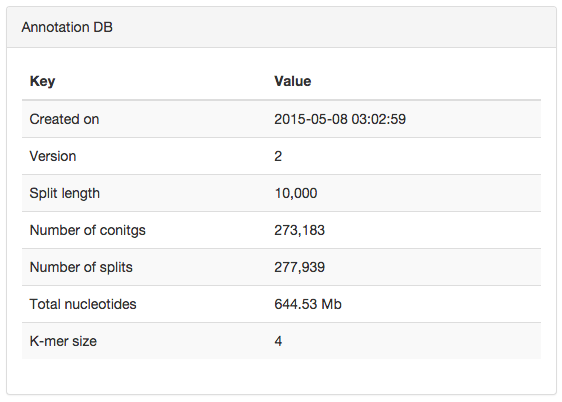
One way to analyze this dataset is to rely on CONCOCT. For instance, you can profile each of your sample with --min-contig-length of 2500, and skip the hierarchical clustering during anvi-merge:
$ anvi-merge */PROFILE.db -c contigs.db -o MERGED_PROFILE --skip-hierarchical-clustering
Although resulting merged profile would be missing all the trees that are used by anvi-interactive, it would contain clustering results from CONCOCT, therefore you would still have your genome bins identified. Which means, you can now run anvi-summarize on the merged data, and take a look at what’s up:
$ anvi-summarize -p MERGED_PROFILE/PROFILE.db -c contigs.contigsdb -C CONCOCT -o MERGED_SUMMARY
Big data is all about trade-offs. Relying on fully-automated genome binning means;
- You will be able to process very large number of contigs and get many many genome bins
- Many of your genome bins are not going to be as pure as you would like them
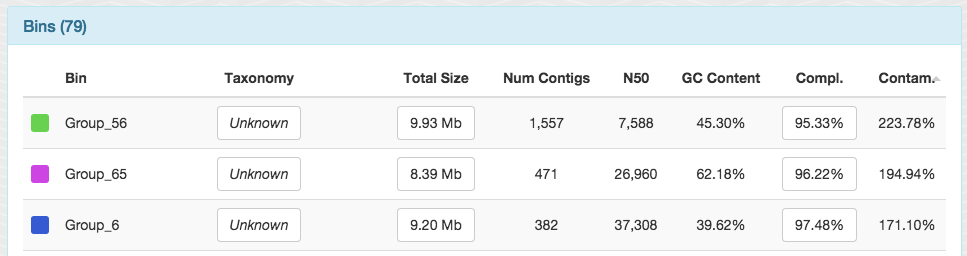
Although CONCOCT largely does great and gives a lot of near-complete genome bins with low contaamination, here is a couple of bins identified by CONCOCT in this dataset with a lot of contamination:
Solution: anvi-refine
Once you are here, you can use anvi-refine to switch to supervised binning and refine them by identifying separate genome bins by hand. Lets take Group_6 as an example:
Fortunately, there aren’t 200,000 contigs in Group_6, which means we can analyze it using the default clustering and visualization workflow of anvi’o. To achieve that we can go back to our terminal and type this command:
I would suggest you to take a backup of your original profile database just in case if things go bad.
$ anvi-refine -p MERGED_PROFILE/PROFILE.db -c contigs.db -C CONCOCT -b Group_6
With this command, anvi’o will take all contigs that were put in Group_6 in the CONCOCT collection, along with the data associated with them stored in the contigs and profile databases, and run a hierarchical clustering on this subset, just to present you with the interactive interface so you can refine this highly contaminated bin.
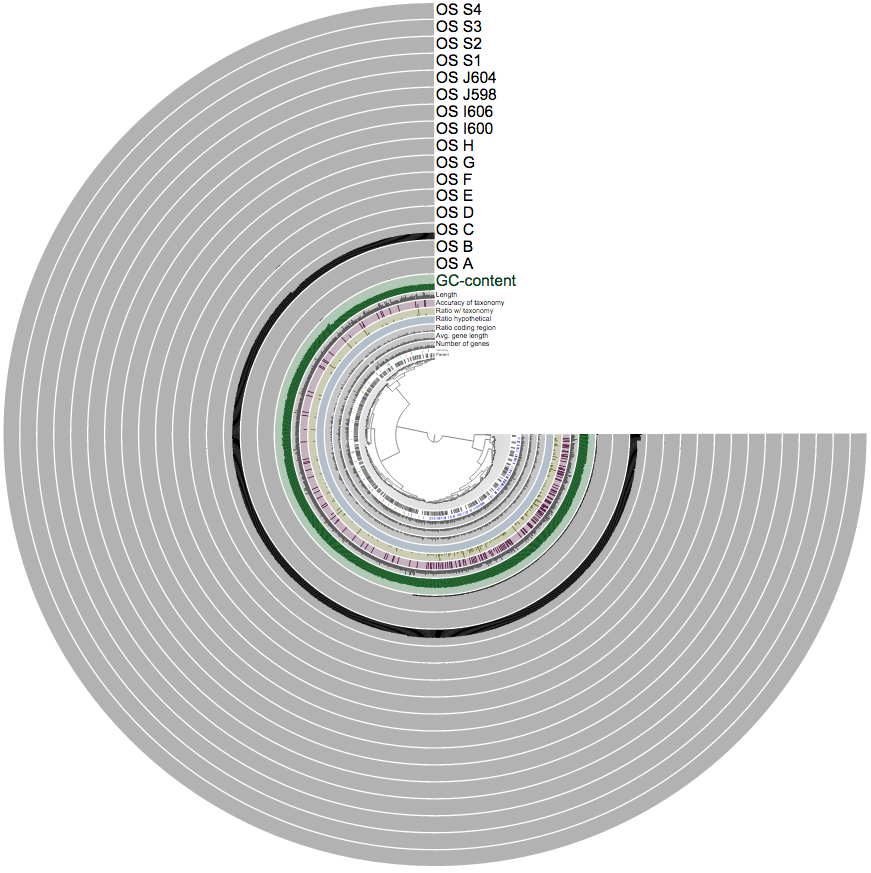
After a short wait while anvi’o gets everything sorted out, you are welcomed with the interactive interface:
This tree shows every contig that was in Group_6, however, they are this time clustered by anvi’o.
You can immediately see why CONCOCT had hard time with them and ended up putting all them together: all these contigs are coming from genomes that are occurred mostly in sample OS C. As CONCOCT relies on differential distribution of genomes across samples, great similarity in the distribution of these genomes makes it very hard to confidently pull them apart. Which is not surprising. However, when we focus on this bin with anvi’o, we can see some tiny differences in distribution:
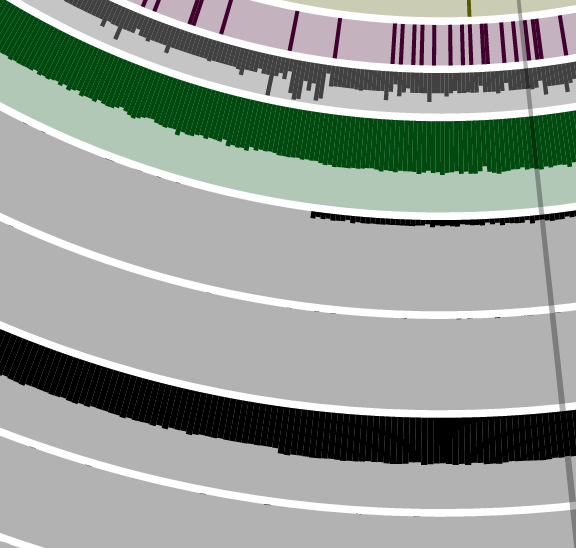
When anvi’o focuses only one CONCOCT group, these tiny differences become large enough to have three, very clear clusters as you can see from the tree:
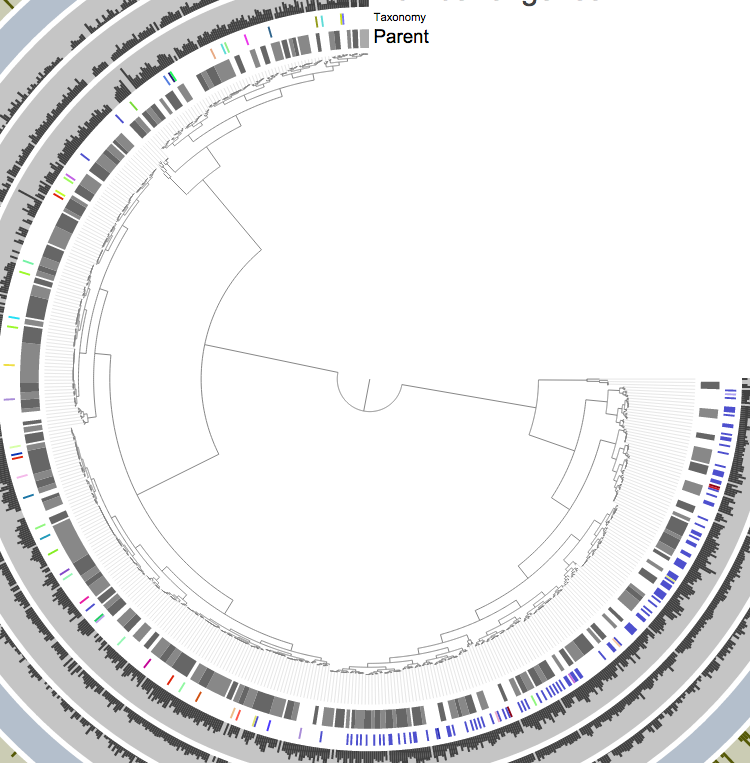
We know when we select all of them, the predicted contamination is about 170%. But if I click those branches to add them into separate groups:
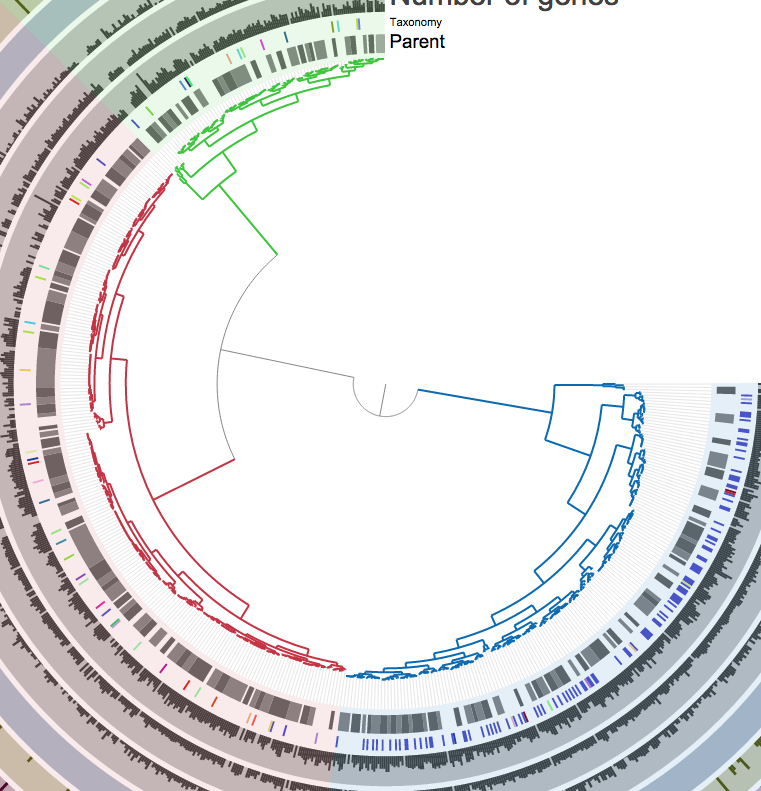
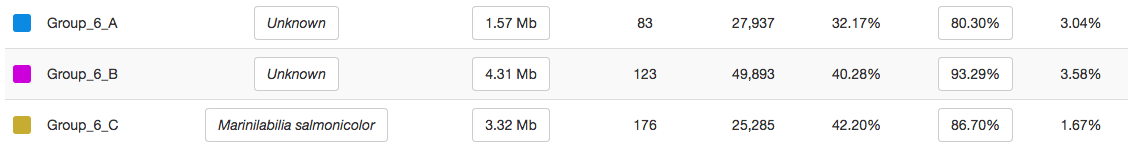
The contamination drops down to much better levels:
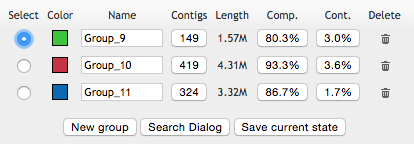
Saving the refined group
When you click ‘save’, the interface will gracefully remove Group_6 from the database, and add these new three groups instead. It is essential to give proper names to newly created groups if you would like to recognize them in the output. Once you are done, of course the next step is to re-run the summary:
$ anvi-summarize -p MERGED_PROFILE/PROFILE.db -c contigs.db -C CONCOCT -o MERGED_SUMMARY
When you look at the output, you will see that this is not there anymore:
And instead, you have this:
Yay! contigs