Table of Contents
- Getting help from anvi’o
- Browse anvi’o programs and artifacts online
- Query anvi’o programs offline
- See all anvi’o programs online
- Explore anvi’o concepts online
- Getting help from the community
- Anvi’o Discord [non-technical]
- Comments on MerenLab.org [non-technical]
- GitHub Issues [technical]
Anvi’o does a lot. And sometimes it can be a little overwhelming. We ourselves often feel overwhelmed by it. That’s why we are writing this post, to remind everyone (including ourselves) how to find the best resources to learn anvi’o and how to find help when we have a problem.
This post is for you if you would like to have a better idea about how to,
- Find anvi’o programs and understand their utility through online and offline means,
- Investigate how anvi’o programs, databases and concepts are connected to each other,
- Ask usage questions and/or discuss scientific ideas and best-practices with the anvi’o community, and,
- Report technical problems with the platform to its developers.
Getting help from anvi’o
The purpose of this section is to introduce you to resources from which you can get online or offline help to better use anvi’o by yourself, without reaching out to another person.
Browse anvi’o programs and artifacts online
Please visit this page for online help for anvi’o programs and artifacts.
Query anvi’o programs offline
You know what you want to do with your data, and you think you can do it with anvi’o. But how do you find an anvi’o program to do it?
Enter anvi-help. A recently implemented program by our very own Evan Kiefl that allows you to query available anvi’o programs using keywords of your choosing. The usage is very straightforward: suppose you want ‘functions’. The following command line will search the keyword ‘functions’ in all anvi’o programs to find those that have anything to do with it:
anvi-help functions
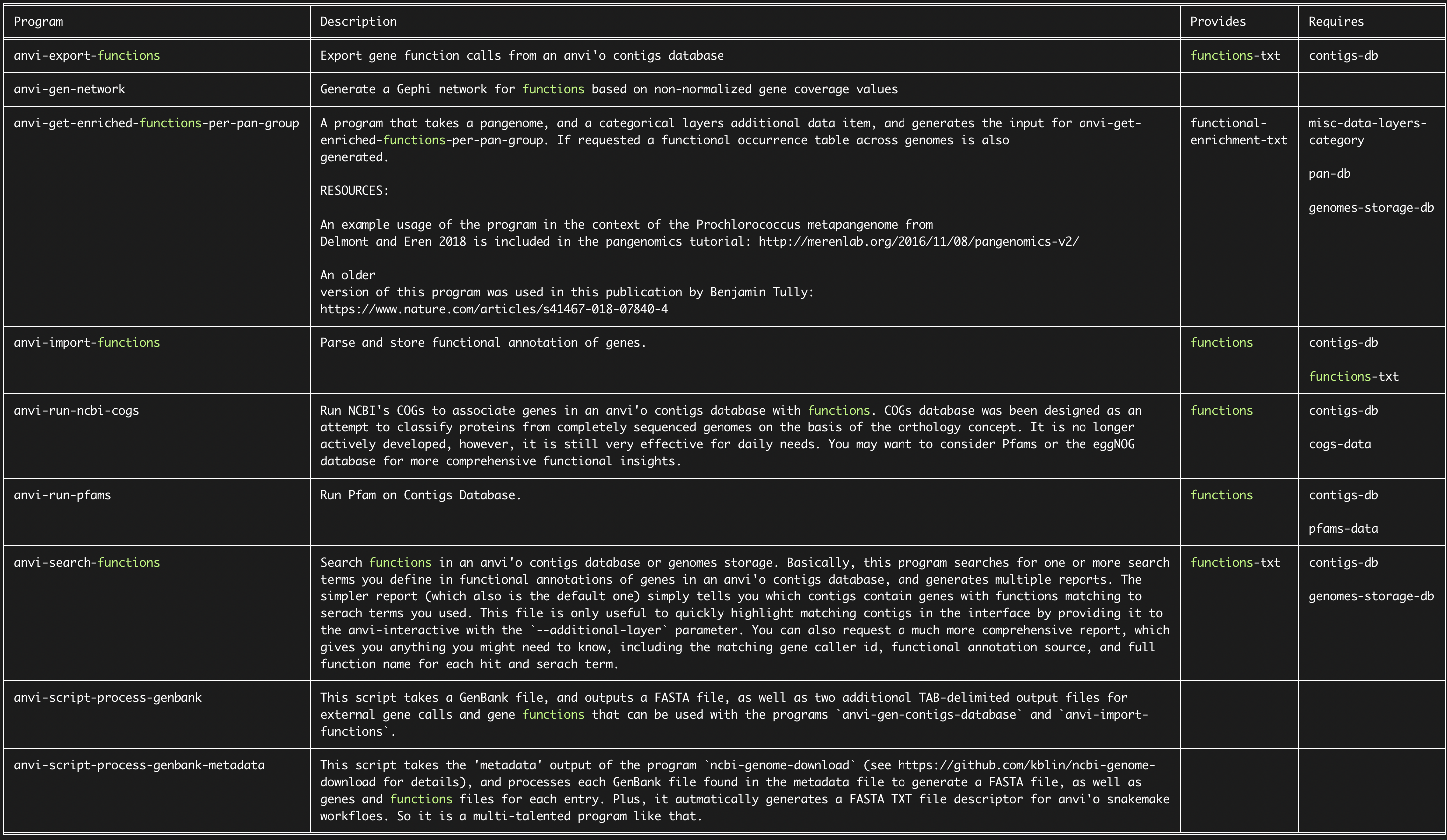
After thinking about itself for a bit, anvi’o will return to you a table that looks like this:
As you can see, there are multiple anvi’o programs related to the keyword “functions”. Let’s assume you are interested using anvi-run-ncbi-cogs since it does provide functions and you like NCBI. From the output, you can see that anvi-run-ncbi-cogs requires contigs-db and cogs-data. Let’s assume you are not familiar with either. Luckily, you can ask anvi-help who provides these requiremements. Let’s start with cogs-data:
anvi-help --provides cogs-data
This indicates all you need to do is to run the program by itself to setup the required COGs data on your computer. How about a contigs database?
anvi-help --provides contigs-database
And there you have it! From three quick searches, you know how to go from the FASTA file of your contigs to a contigs database with COG functions using three anvi’o programs. If you were to ask for fun whether anything in anvi’o provides FASTA files, you could search for that as well,
anvi-help --provides fasta
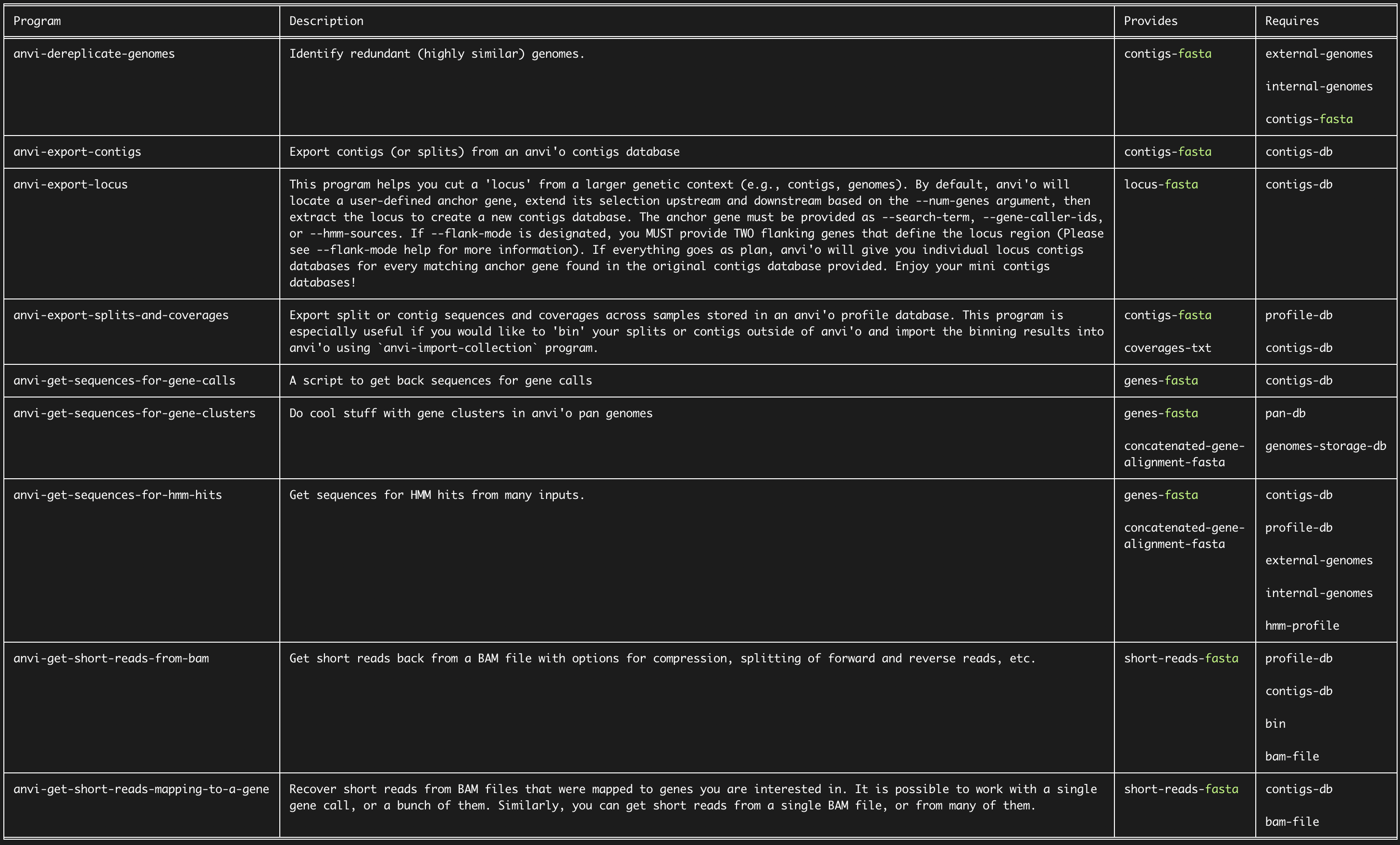
And perhaps learn some things that you didn’t think you needed before:
See all anvi’o programs online
The vignette is the place to go for a comprehensive list of information on all programs in the latest stable release of anvi’o. It shows much of the same information that anvi-help shows and individual program help pages (-h) would give you, sometimes with extra tidbits of explanation. Googling an anvi’o program usually directs you to the vignette page. But if you are one of those adventurous folks who is using the development version of anvi’o (good for you!), some of the newer programs may not be on this page yet.
Explore anvi’o concepts online
Anvi’o is a web of interconnected concepts, data, and programs. The program anvi-help traverses what anvi’o programs know about themselves to help you find your way starting with a keyword. An alternative way to do it is to browse interactively how everything in anvi’o is connected as a network. We have a nice visualization of this network that is useful for exploring what anvi’o can do and what it needs from you to do it.
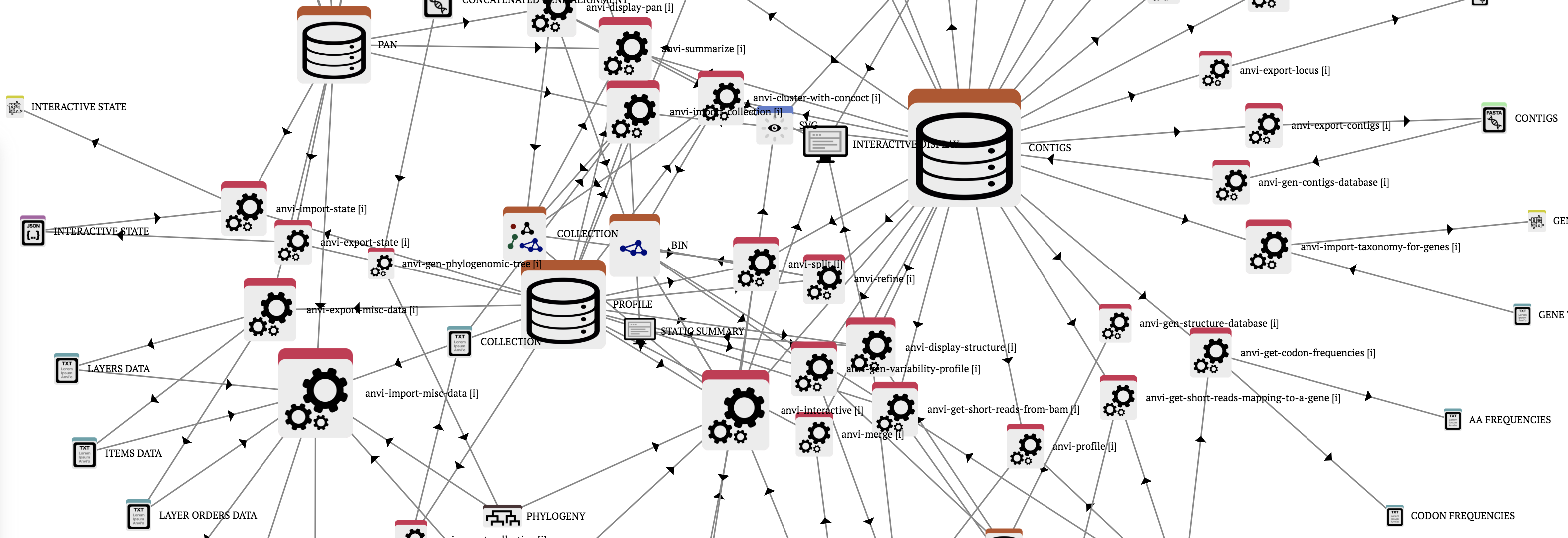
At first glance, the network may look complicated:
But you can click on any one icon to learn more about how it fits into the rest of the anvi’o universe. This helps you to quckly learn more about what you can do with a contigs database, for instance. For that, you would click on the contigs-db icon in the network above, and peruse the vast web of anvi’o programs that use or generate these databases.
Getting help from the community
The purpose of this section is to introduce you to online resources from which you can get help from other people to solve your conceptual, usage, or technical problems with anvi’o, or discuss best-practices for microbial ‘omics in general.
Anvi’o has a friendly user community that is growing and there are a few different ways to reach out to the developers or users in this community depending on your needs. Before listing ways to communicate with us, let’s define what is a technical and a non-technical problem, because each of these require different communication channels.
Definition of non-technical problems
You have a non-technical problem if you are,
- Getting a nicely formatted error message from anvi’o, and/or anvi’o is asking you to contact someone.
- Having trouble running or installing anvi’o.
- Not sure how to do a particular analysis in anvi’o, and cannot find any hints or documentation about it in our online resources.
- Not sure how to set parameters of an anvi’o program to accomplish your particular goal, or have any other such question about more nuanced usage of anvi’o.
- In need of a new feature that is not in anvi’o.
- Confused by any part of our online documentation.
In these cases you can use any of the non-technical communication channels below. But in all cases please makes sure you start your message with a short description of your system (i.e., the output of anvi-self-test -v would be a good information to provide).
Definition of technical problems
You have a technical problem if you are,
- Getting a Python traceback from anvi’o (i.e., an ugly bunch of lines of text as anvi’o crashes on you).
That’s it. This is the only time you have a technical problem that requires you to contact the developers.
The following resources are at your disposal to seek help:
Anvi’o Discord [non-technical]
The anvi’o Discord channel (for which you can always find an up-to-date link on https://anvio.org) is the most effective way to get help from the community. While the conversations on Disord will not be cashed by search engines and will only be accessible to those who join the Discord server. But it is free to join, so that’s that. Click the button below to get your invitation:
We enjoy discussing concepts of microbial ‘omics, so scientific discussions beyond anvi’o are most welcome there.
Comments on MerenLab.org [non-technical]
Most of our blog posts, tutorials, and other web pages have a comments section at the bottom where you can ask your questions (or give your feedback) on issues related to the topic discussed in the post/tutorial/page. This is often very useful to the rest of the community, as it collects relevant discussions right there where other people will go to look for that information.
GitHub Issues [technical]
If you are having a technical problem, please submit an issue on GitHub to contact developers.
The only time it is appropriate to use GitHub is if you are having a technical issue, or a developer requested you to submit an issue for a non-technical concern.
When you are submitting a GitHub issue, it is extremely critical for you to read the issue template that appears when you click “New Issue”, and remove the instructions once you have read them all before submission.
Educational resources
We try our best to keep our online documentation up-to-date. Our best is not always enough, but most of the time you should be able to find plenty of information right here on our website. This section gives a brief overview of the different kinds of learning resources we provide for beginners.
The ‘omics vocabulary
In anvi’o programs and most of the documents referenced below, you will find a whole host of terms of microbial ‘omics that are specific to our field (such as ‘contig’) or specific to anvi’o (such as ‘contigs database’). While familiarity with the terms is essential, they can be initially confusing and seemingly impenetrable. Which is why we have started developing a dedicated vocabulary page to help you get started.
Our vocabulary page is in its infancy. If someting is missing, or if you do not agree with some of the definitions, we would be happy to hear from you.
Tutorials
Anvi’o includes various sets of programs that are meant to be used together to accomplish a larger analysis task. We often write tutorials to explain these common use cases. These are step-by-step instructions on how to get from your initial data (FASTA files, BAM files, etc) to visualizations of biological insights, in anvi’o. They can be accessed from the “Tutorials” pull-down menu on each page of the anvi’o section of our website.
Tutorials under the ‘More Abstract’ category are generic recipes. They focus on the commands that should be run to get from A to B, without discussion of the finer details of data interpretation and visualization at each step. These are better suited for people who know what they want to do and generally how they could do it, but need to figure out how to perform their analysis in anvi’o.
In contrast, tutorials under the ‘More Practical’ category are geared towards more comprehensive instruction, using a specific dataset as an example. The datasets are downloadable, allowing you to follow along by running the commands on your own computer and comparing the output you get to that shown in the tutorial. These are suitable for people who know what they want to do but not how to do it. Caveats, expert tips, and visualization strategies that go along with each step are often discussed, making these tutorials excellent for those who want to understand each step of the process and need some context to get it.
Please note that tutorials are often written around the time when their core programs are first published. This (unfortunately) often makes them a snapshot of how to do an analysis with a particular version of anvi’o, rather than an evolving document. We update them occasionally, but finer details may escape our attention. The consequences of this are that specific commands might not work for you, depending on how much anvi’o has been updated since the tutorial was written. If that happens, don’t worry! The overall analysis workflow usually remains the same, and there are plenty of resources to help you figure out how to modify the command. Please feel free to send us a comment, email, or slack message about it - we can help you out, and then update our tutorial so that other people don’t struggle with the same issue :)
Workshops
We periodically give workshops for hands-on training on anvi’o (and we would be thrilled to make your acquaintance if you can make it to one). This page keeps a list of past and upcoming workshops. Usually anvi’o workshops are quite fun, and the only thing you need to bring is your laptop computer.
A course on microbial ‘omics
We are always searching for ways to make our field of science more accessible to everyone. For those who want to learn more about microbial ‘omics in general, you can check out Meren’s classroom resources on the topic, and feel free to get in touch if you are an educator and wishes to use anvi’o for your course.
Please let us know if there is a way to improve anything on this page.