Table of Contents
Our daily workflow with anvi’o usually requires us to use some server-side computing for profiling metagenomic samples, followed by the visualization of merged profiles through the interactive interface on our laptops. The second step requires us to download large amounts of data, which can be far from what is feasible in certain situations.
Fortunately, you can connect to a server by creating an SSH tunnel with local and remote port forwarding, and run the interactive interface on the server to access it from your laptop.
For this, you should connect to your server in a specific way:
local ~ $ ssh -L 8080:localhost:8080 meren@server.university.eduIf you care, the translation of this line is this: “forward any connection request goes to my local port 8080, to the port 8080 on localhost of the server server.university.edu”. Of course nothing is listening to the port 8080 on the server at this moment, but we will tell anvi’o to serve from there. For this, go to your data directory:
server ~ $ cd my_data/
server ~/my_data $ ls
CONTIGS.db MERGED/ SAMPLES.db (...)And run anvi-interactive:
server ~/my_data $ anvi-interactive -p MERGED/PROFILE.db -c CONTIGS.db -s SAMPLES.db --server-only -P 8080
Contigs DB .........................: Initialized: CONTIGS.db (v. 3)
Auxiliary Data .....................: Found: MERGED/AUXILIARY-DATA.db (v. 1)
Profile DB .........................: Initialized: MERGED/PROFILE.db (v. 6)
* The server is now listening the port number "8080". When you are finished, press
CTRL+C to terminate the server.Notice the --server-only and -P 8080 flags. If port 8080 is not available, you can re-connect to the server with a differnt forwarding request via SSH, and try that port number for anvi’o to serve.
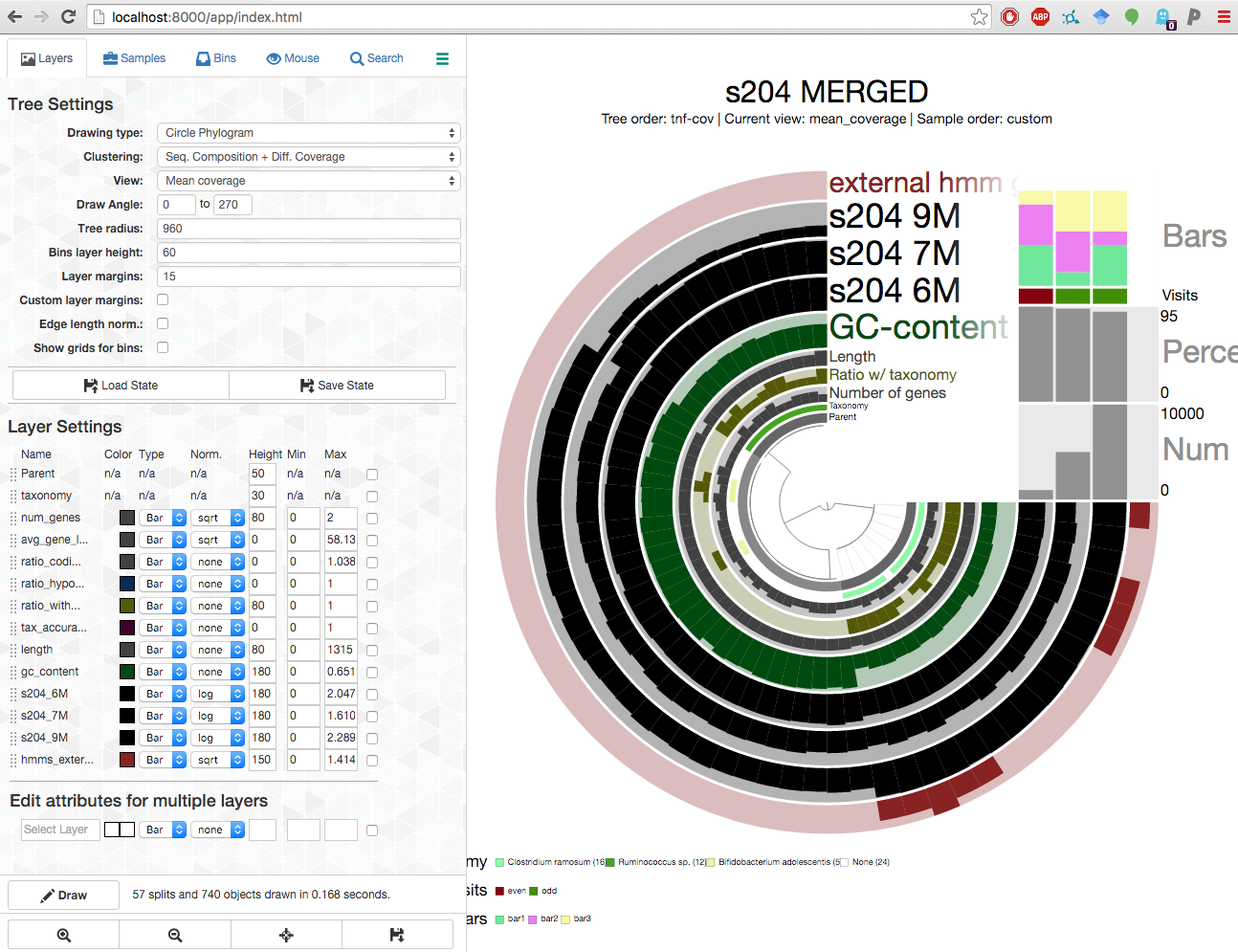
All good on the server side. Now you can start a browser on your laptop computer, and type the address http://localhost:8080 to stream your results from the server to your browser like a pro: