Table of Contents
- Thanks
- Introduction
- Setting up anvi’o SCG taxonomy
- Populating contigs db with SCG taxonomy
- Estimating taxonomy in the terminal
- Contigs db for a single genome
- Contigs db for a single metagenome
- Contigs db + profile db
- Contigs db + profile db + collection
- Many contigs dbs for many metagenomes
- Estimating taxonomy in the interface
- Final words and some warnings
The latest version of anvi’o is v7.1. See the release notes.
The shortened URL for this page is https://merenlab.org/scg-taxonomy
Although anvi’o SCG taxonomy has been available since anvi’o v6, this tutorial is designed for v6.2 and later versions of anvi’o. You can identify which version you have on your computer by typing anvi-self-test --version in your terminal. If you are using anvi’o v6.1 some commands may not work as expected.
With the anvi’o scg-taxonomy workflow you can learn,
- Taxonomic affiliation of a single genome (you have a contigs db)
- Taxonomic profile of a single metagenome (you have a contigs db)
- Relative abundances of taxa in a given metagenome (you have a contigs db and a single profile db)
- Taxonomic profiles of genome bins described in an anvi’o collection (you have a contigs db, single or merged profile db, and a collection (i.e., binning results))
- Taxonomic profiles across metagenomes (you have many contigs dbs and optionally single profile dbs)
- and perhaps most importantly, real-time taxonomy estimation of a new bin while working with anvi’o interactive interface!
If you find a mistake on this page or would you like to update something in it, please feel free to edit its source by clicking the edit button at the top-right corner (which you will see if you are logged in to GitHub) 😇
Thanks
The new anvi’o scg-taxonomy module described on this page stands on more than 2,500 lines of code that is fully integrated to the rest of the anvi’o platform. This effort is partially a result of a pilot award from the FACCTS (France and Chicago Collaborating in The Sciences) program, which supported the travel expenses of Quentin Clayssen from France to Chicago, who put the initial effort to investigate the feasibility of this idea. I would also like to thank Alon Shaiber for his excellent suggestions that influenced the speed of this algorithm, and Özcan Esen for helping with the integration of this module to the anvi’o interface.
Introduction
Often when we deal with genomes we need some sort of insight into taxonomy. This becomes an especially critical need when you reconstruct genomes from metagenomes, as you want to know where these new genomes fit in the tree of life rapidly. So far anvi’o did not offer a rapid solution for that, but starting with the v6 we have a solution for that. Everything is new, so probably there are going to be hiccups before we manage to stabilize things.
The new anvi’o scg-taxonomy worklow uses the taxonomy determined by the The Genome Taxonomy Database (GTDB, @ace_gtdb), an effort led by scientists who are primarily affiliated with the Australian Centre for Ecogenomics. And the workflow searches new seqeunces in GTDB databases using DIAMOND, a fast alternative to the NCBI’s BLAST, developed and maintained by the brilliant Benjamin Buchfink, who is a bioinformatician at the MPI for Developmental Biology in Germany. We thank them sincerely.
If you use anvi’o scg-taxonomy module, please don’t forget to cite the initial work that introduced this resource (doi:10.1038/nbt.4229), and DIAMOND (doi:10.1038/nmeth.3176).
Please note that there is another tool from the developers of the GTDB, GTDB-Tk, to assign taxonomy to genomes.
In a nutshell, anvi’o (1) downloads both single-copy core genes (SCGs) and the taxonomy of the genomes - as defined by the GTDB - from which these genes are coming from, (2) bulids search databases for a subset of these SCGs (depending on whether they are in the Bacterial and Archaeal collection of SCGs anvi’o recognizes and based on other heuristics), (3) searches SCGs from every new genome in the anvi’o ecosystem in these databases and creates consensus taxonomy for each genome.
The following sections will cover the three major steps for anvi’o scg-taxonomy: (1) setting things up, (2) populating contigs database with taxonomy, and (3) running taxonomy estimations. The text will cover how to run anvi’o scg-taxonomy both from the terminal environment and through the graphical user interface.
For the sake of reproducibility, here I will use the infant gut dataset that we use to demonstrate many things anvi’o. If you would like to follow this tutorial not with your own datasets but with the infant gut dataset, you can simply click here first, follow the download instructions, and come back here once the text tells you to go back.
Setting up anvi’o SCG taxonomy
This is something you will do only once after installing anvi’o v6.
SCG-TAXONOMY WILL NOT WORK unless you have diamond version 0.9.14 installed on your system. Please first run diamond --version to make sure you have the right version. If not, install it first. If you are in a conda environment, you can simply run conda install diamond=0.9.14.
A fresh installation of anvi’o will have all the FASTA files that are going to be used for scg-taxonomy, but not the search databases that must be generated from them. We are not shipping the databases because they need to be built with the DIAMOND installed on your computer to make sure we are not running into various incompatibilty problems.
To setup your SCG taxonomy databases, run the following program, which will not take longer than a minute:
anvi-setup-scg-taxonomy
This program is called anvi-setup-scg-databases in anvi’o v6.2 or earlier versions.
Please see the help menu for every program even if the text does not ask you to do it. There are many parameters for every anvi’o program so you can tweak things depending on your system and needs.
When I run this on my computer, this is the output I get:
(anvio-master) (base) meren ~ $ anvi-setup-scg-taxonomy
WARNING
===============================================
Please remember that the data anvi'o uses for SCG taxonomy is a courtesy of The
Genome Taxonomy Database (GTDB), an initiative to establish a standardised
microbial taxonomy based on genome phylogeny, primarly funded by tax payers in
Australia. Please don't forget to cite the original work, doi:10.1038/nbt.4229
by Parks et al to explicitly mention the source of databases anvi'o relies upon
to estimate genome level taxonomy. If you are not sure how it should look like
in your methods sections, anvi'o developers will be happy to help you if you
can't find any published example to get inspiration.
WARNING
===============================================
Anvi'o found your FASTA files in place, but not the databases. Now it will
generate all the search databases using the existing FASTA files.
* Every FASTA is now turned into a fancy search database. It means you are now
allowed to run `anvi-run-scg-taxonomy` on anvi'o contigs databases.
If you don’t have permission to write to the part of the disk anvi’o is installed on your system, you should ask your system administrator to run this command for you.
The command above will only create the databases without downloading original files. In some rare cases you may have to re-download everything and start with a clean slate. If you think this is what you need, try this command (but you should run this only if you have a problem):
anvi-setup-scg-taxonomy --reset
Populating contigs db with SCG taxonomy
This is something you will do once for every contigs database you wish to work with.
For anvi’o scg-taxonomy to work, you need to run the following program on your contigs database:
anvi-run-scg-taxonomy -c CONTIGS.db
If you have multiple CPUs and memory you may want to use --num-parallel-processes and --num-threads parameters to speed things up. Please refer to the help menu to find out what they actually mean. Just to give you an idea, running this program on the infant gut dataset without any parallelization takes 50 seconds. When I run the same command with --num-parallel-processes 3 and --num-threads 3, it takes 10 seconds. Just so you know.
Here is a crude list of what is going on behind the scenes when you run this program:
- Anvi’o finds all the 22 single-copy core genes it uses for scg-taxonomy in a given contigs database,
- Searches them against the corresponding databases from the GTDB, and,
- Chooses the best hit among all the hits, and stores them in a table in the contigs database.
If you run the program with the --debug flag you can see all the underlying taxonomy strings that lead to the consensus. Beware, though, your terminal will be overwhelmed by some text :)
Estimating taxonomy in the terminal
There are many ways to estimate taxonomy, but all of them are going to use the program anvi-estimate-scg-taxonomy. Following sections will demonstrate multiple uses of this tool.
Contigs db for a single genome
If you have an anvi’o contigs for a single genome, you can simply run this command:
anvi-estimate-scg-taxonomy -c CONTIGS.db
For instance, when I run this on an Enterococcus faecalis genome I have downloaded from the NCBI, this is what I get:
anvi-estimate-scg-taxonomy -c Enterococcus_faecalis_6240.db
Contigs DB ...................................: Enterococcus_faecalis_6240.db
Profile DB ...................................: None
Metagenome mode ..............................: False
Estimated taxonomy for "Enterococcus_faecalis_6240"
===============================================
╒════════════════════════════╤══════════════╤═══════════════════╤════════════════════════════════════════════════════════════════════════════════════════════════════════════╕
│ │ total_scgs │ supporting_scgs │ taxonomy │
╞════════════════════════════╪══════════════╪═══════════════════╪════════════════════════════════════════════════════════════════════════════════════════════════════════════╡
│ Enterococcus_faecalis_6240 │ 21 │ 20 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
╘════════════════════════════╧══════════════╧═══════════════════╧════════════════════════════════════════════════════════════════════════════════════════════════════════════╛
Total SCGs and Supporting SCGs columns indicate the number of SCGs found in this genome that were suitable for taxonomy estimation, and the number of SCGs that matched to the consensus taxonomy. You may be curious why there is 21 SCGs but only 20 are supporting the consensus. To see a more detailed view of the SCGs and their relationship with the consensus taxonomy, you can run the same command with --debug flag:
anvi-estimate-scg-taxonomy -c Enterococcus_faecalis_6240.db --debug
Contigs DB ...................................: Enterococcus_faecalis_6240.db
Profile DB ...................................: None
Metagenome mode ..............................: False
* A total of 21 single-copy core genes with taxonomic affiliations were
successfully initialized from the contigs database 🎉 Following shows the
frequency of these SCGs: Ribosomal_S2 (1), Ribosomal_S3_C (1), Ribosomal_S6 (1),
Ribosomal_S7 (1), Ribosomal_S8 (1), Ribosomal_S9 (1), Ribosomal_S11 (1),
Ribosomal_S20p (1), Ribosomal_L1 (1), Ribosomal_L2 (1), Ribosomal_L3 (1),
Ribosomal_L4 (1), Ribosomal_L6 (1), Ribosomal_L9_C (1), Ribosomal_L13 (1),
Ribosomal_L16 (1), Ribosomal_L20 (1), Ribosomal_L21p (1), Ribosomal_L22 (1),
ribosomal_L24 (1), Ribosomal_L27A (1), Ribosomal_L17 (0).
Hits for Enterococcus_faecalis_6240
===============================================
╒════════════════╤════════╤══════════╤════════════════════════════════════════════════════════════════════════════════════════════════════════════╕
│ SCG │ gene │ pct id │ taxonomy │
╞════════════════╪════════╪══════════╪════════════════════════════════════════════════════════════════════════════════════════════════════════════╡
│ Ribosomal_S6 │ 2178 │ 97.9 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L27A │ 2372 │ 99.3 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L1 │ 1541 │ 100.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L9_C │ 2182 │ 97.4 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S11 │ 2378 │ 100.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S9 │ 2067 │ 100.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L13 │ 2068 │ 99.3 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L21p │ 150 │ 99.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L20 │ 2970 │ 99.1 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S20p │ 1318 │ 98.8 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S7 │ 2346 │ 100.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L3 │ 2352 │ 99.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L4 │ 2353 │ 100.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L2 │ 2355 │ 99.6 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L22 │ 2357 │ 100.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S3_C │ 2358 │ 99.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L16 │ 2359 │ 100.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ ribosomal_L24 │ 2363 │ 98.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S2 │ 1276 │ 100.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S8 │ 2366 │ 99.2 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_L6 │ 2367 │ 100.0 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────┼────────┼──────────┼────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ CONSENSUS │ -- │ -- │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
╘════════════════╧════════╧══════════╧════════════════════════════════════════════════════════════════════════════════════════════════════════════╛
Estimated taxonomy for "Enterococcus_faecalis_6240"
===============================================
╒════════════════════════════╤══════════════╤═══════════════════╤════════════════════════════════════════════════════════════════════════════════════════════════════════════╕
│ │ total_scgs │ supporting_scgs │ taxonomy │
╞════════════════════════════╪══════════════╪═══════════════════╪════════════════════════════════════════════════════════════════════════════════════════════════════════════╡
│ Enterococcus_faecalis_6240 │ 21 │ 20 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
╘════════════════════════════╧══════════════╧═══════════════════╧════════════════════════════════════════════════════════════════════════════════════════════════════════════╛
From this output we learn that Ribosomal_L16 only resolves to the genus level, hence not matching perfectly to the consensus. Fine. We can live with that.
Contigs db for a single metagenome
If you are working with a contigs database that represents a metagenome, anvi’o will complaining that there is too much redundancy of SCGs for this to be a single genome. You can avoid that by using the flag --metagenome-mode. For this example, let’s use the famous Infant Gut Data (if you would like to follow these steps, you can obtain this dataset and set it up on your computer following the instructions here).
The Infant Gut Data (IGD) contains a contigs database that represents a metagenomic assembly, so it is appropriate to investigate taxa occurring in this metagenome through this command:
anvi-estimate-scg-taxonomy -c CONTIGS.db \
--metagenome-mode
First let’s take a look at the output, and then discuss what is happening behind the scenes:
Contigs DB ...................................: CONTIGS.db
Profile DB ...................................: None
Metagenome mode ..............................: True
SCG for metagenome ...........................: None
* A total of 171 single-copy core genes with taxonomic affiliations were
successfully initialized from the contigs database 🎉 Following shows the
frequency of these SCGs: Ribosomal_S6 (10), Ribosomal_S8 (10), Ribosomal_L27A
(10), Ribosomal_S3_C (9), Ribosomal_S9 (9), Ribosomal_L4 (9), Ribosomal_L6 (9),
Ribosomal_L9_C (9), Ribosomal_L16 (9), ribosomal_L24 (9), Ribosomal_L1 (8),
Ribosomal_S20p (7), Ribosomal_L2 (7), Ribosomal_L13 (7), Ribosomal_L17 (7),
Ribosomal_L22 (7), Ribosomal_S2 (6), Ribosomal_S11 (6), Ribosomal_L3 (6),
Ribosomal_L20 (6), Ribosomal_L21p (6), Ribosomal_S7 (5).
WARNING
===============================================
Anvi'o automatically set 'Ribosomal_S6' to be THE single-copy core gene to
survey your metagenome for its taxonomic composition. If you are not happy with
that, you could change it with the parameter `--scg-name-for-metagenome-mode`.
Taxa in metagenome "Infant Gut Contigs from Sharon et al."
===============================================
╒════════════════════╤════════════════════╤══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╕
│ │ percent_identity │ taxonomy │
╞════════════════════╪════════════════════╪══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╡
│ Ribosomal_S6_11655 │ 98.9 │ Bacteria / Firmicutes / Bacilli / Staphylococcales / Staphylococcaceae / Staphylococcus / Staphylococcus epidermidis │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S6_12163 │ 98.9 │ Bacteria / Firmicutes / Bacilli / Staphylococcales / Staphylococcaceae / Staphylococcus / Staphylococcus hominis │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S6_15200 │ 98.9 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Lactobacillaceae / Leuconostoc / Leuconostoc citreum │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S6_25880 │ 98.9 │ Bacteria / Firmicutes / Clostridia / Tissierellales / Helcococcaceae / Finegoldia / Finegoldia magna │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S6_2915 │ 97.9 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S6_29818 │ 98.9 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Streptococcaceae / Streptococcus / Streptococcus oralis │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S6_30904 │ 92.5 │ Bacteria / Firmicutes / Clostridia / Tissierellales / Helcococcaceae / Anaerococcus / Anaerococcus sp002359915 │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S6_4484 │ 98.9 │ Bacteria / Firmicutes / Clostridia / Tissierellales / Peptoniphilaceae / Peptoniphilus / Peptoniphilus rhinitidis │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S6_6421 │ 100 │ Bacteria / Actinobacteriota / Actinobacteria / Propionibacteriales / Propionibacteriaceae / Cutibacterium / Cutibacterium avidum │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S6_7660 │ 98.9 │ Bacteria / Firmicutes / Bacilli / Staphylococcales / Staphylococcaceae / Staphylococcus / │
╘════════════════════╧════════════════════╧══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╛
Here is a simplified list of what is happening when you run this command:
- Anvi’o counts the occurrence of each of the 22 single-copy core gene names in this contigs database,
- Chooses the most frequent one to use for taxonomy estimation, and,
- Generates an output.
All these fancy terminal outputs can be stored in cleaner, flat TAB-delimited files using --output-file parameter.
Please note that anvi’o uses all SCGs and predicts a consensus taxonomy in genome mode, but chooses the most frequent SCG to predict taxa in metagenome mode. The discrepancy between genome and metagenome mode comes from the fact that we need to focus on a single SCG to get a reasonable taxonomy profile from a metagenome. You can get a flat text file that shows frequencies of each SCG by using the parameter --report-scg-frequencies. Please remember that one of the most significant shortcomings of this workflow is that you will not find taxonomic signal for those populations that did not assemble at all.
Let’s go back to the output of the previous command. One of the first messages you see in that output is this, where anvi’o reports the frequencies of SCGs:
* A total of 171 single-copy core genes with taxonomic affiliations were
successfully initialized from the contigs database 🎉 Following shows the
frequency of these SCGs: Ribosomal_S6 (10), Ribosomal_S8 (10), Ribosomal_L27A
(10), Ribosomal_S3_C (9), Ribosomal_S9 (9), Ribosomal_L6 (9), Ribosomal_L9_C
(9), Ribosomal_L16 (9), ribosomal_L24 (9), Ribosomal_L1 (8), Ribosomal_L2 (8),
Ribosomal_L4 (8), Ribosomal_S20p (7), Ribosomal_L13 (7), Ribosomal_L17 (7),
Ribosomal_L22 (7), Ribosomal_S2 (6), Ribosomal_S11 (6), Ribosomal_L3 (6),
Ribosomal_L20 (6), Ribosomal_L21p (6), Ribosomal_S7 (5).
Because there are 10 Ribosomal_S6 genes, anvi’o automatically used this SCG to estimate taxonomy. For instance, Ribosomal_S9 occurs 9 times instead of 10, we can ask anvi’o to use that one instead the following way:
anvi-estimate-scg-taxonomy -c CONTIGS.db \
--metagenome-mode \
--scg-name Ribosomal_S9
I get a similar picture of the same data, but of course I am missing one of the entries, reminding you the major limitation of this strategy:
════════════════════╤════════════════════╤══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╕
│ │ percent_identity │ taxonomy │
╞════════════════════╪════════════════════╪══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╡
│ Ribosomal_S9_10246 │ 100 │ Bacteria / Firmicutes / Clostridia / Tissierellales / Peptoniphilaceae / Peptoniphilus / Peptoniphilus rhinitidis │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S9_14976 │ 100 │ Bacteria / Firmicutes / Bacilli / Staphylococcales / Staphylococcaceae / Staphylococcus / Staphylococcus hominis │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S9_16436 │ 100 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Lactobacillaceae / Leuconostoc / Leuconostoc citreum │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S9_2307 │ 100 │ Bacteria / Firmicutes / Bacilli / Staphylococcales / Staphylococcaceae / Staphylococcus / │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S9_25181 │ 100 │ Bacteria / Firmicutes / Clostridia / Tissierellales / Helcococcaceae / Finegoldia / Finegoldia magna │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S9_26102 │ 98.3 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Streptococcaceae / Streptococcus / │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S9_30207 │ 100 │ Bacteria / Firmicutes / Bacilli / Staphylococcales / Staphylococcaceae / Staphylococcus / │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S9_3532 │ 100 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├────────────────────┼────────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Ribosomal_S9_9744 │ 99.2 │ Bacteria / Actinobacteriota / Actinobacteria / Propionibacteriales / Propionibacteriaceae / Cutibacterium / Cutibacterium avidum │
╘════════════════════╧════════════════════╧══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╛
These may not be conclusive taxonomic insights, but it’s still very useful to have a rapid idea about what you have. After all what is a conclusive taxonomic insight anyway?
Contigs db + profile db
Inferring taxonomic membership is one thing, but how about relative abundances of taxa?
If you have a contigs database for your genomes or metagenomes AS WELL AS a single or merged profile database for the same project, you can actually estimate the coverages of those SCGs to which you are assigning taxonomy. To do that, let’s take a look at the output fo the following command on the same dataset:
anvi-estimate-scg-taxonomy -c CONTIGS.db \
-p PROFILE.db \
--metagenome-mode \
--compute-scg-coverages
Tadaa:
Taxa in metagenome "Infant Gut Contigs from Sharon et al."
===============================================
╒════════════════════╤════════════════════╤════════════════════════════════╤═══════════╤═══════════╤══════════╤═══════════╤═══════════╤══════════════╕
│ │ percent_identity │ taxonomy │ DAY_15A │ DAY_15B │ DAY_16 │ DAY_17A │ DAY_17B │ ... 6 more │
╞════════════════════╪════════════════════╪════════════════════════════════╪═══════════╪═══════════╪══════════╪═══════════╪═══════════╪══════════════╡
│ Ribosomal_S6_2915 │ 97.9 │ (s) Enterococcus faecalis │ 372.512 │ 699.853 │ 663.241 │ 186.34 │ 1149.6 │ ... 6 more │
├────────────────────┼────────────────────┼────────────────────────────────┼───────────┼───────────┼──────────┼───────────┼───────────┼──────────────┤
│ Ribosomal_S6_11655 │ 98.9 │ (s) Staphylococcus epidermidis │ 112.694 │ 172.478 │ 147.304 │ 23.3901 │ 140.769 │ ... 6 more │
├────────────────────┼────────────────────┼────────────────────────────────┼───────────┼───────────┼──────────┼───────────┼───────────┼──────────────┤
│ Ribosomal_S6_4484 │ 98.9 │ (s) Peptoniphilus rhinitidis │ 0 │ 0 │ 0 │ 0 │ 0 │ ... 6 more │
├────────────────────┼────────────────────┼────────────────────────────────┼───────────┼───────────┼──────────┼───────────┼───────────┼──────────────┤
│ Ribosomal_S6_7660 │ 98.9 │ (g) Staphylococcus │ 0 │ 0 │ 0 │ 0 │ 0 │ ... 6 more │
├────────────────────┼────────────────────┼────────────────────────────────┼───────────┼───────────┼──────────┼───────────┼───────────┼──────────────┤
│ Ribosomal_S6_6421 │ 100 │ (s) Cutibacterium avidum │ 17.8935 │ 5.87368 │ 4.79909 │ 0 │ 0 │ ... 6 more │
├────────────────────┼────────────────────┼────────────────────────────────┼───────────┼───────────┼──────────┼───────────┼───────────┼──────────────┤
│ Ribosomal_S6_12163 │ 98.9 │ (s) Staphylococcus hominis │ 2.39322 │ 22.5447 │ 13.2806 │ 0 │ 9.94853 │ ... 6 more │
├────────────────────┼────────────────────┼────────────────────────────────┼───────────┼───────────┼──────────┼───────────┼───────────┼──────────────┤
│ Ribosomal_S6_15200 │ 98.9 │ (s) Leuconostoc citreum │ 0 │ 0 │ 1.86532 │ 0 │ 1.6936 │ ... 6 more │
├────────────────────┼────────────────────┼────────────────────────────────┼───────────┼───────────┼──────────┼───────────┼───────────┼──────────────┤
│ Ribosomal_S6_29818 │ 98.9 │ (s) Streptococcus oralis │ 0 │ 0 │ 0 │ 0 │ 0 │ ... 6 more │
├────────────────────┼────────────────────┼────────────────────────────────┼───────────┼───────────┼──────────┼───────────┼───────────┼──────────────┤
│ Ribosomal_S6_25880 │ 98.9 │ (s) Finegoldia magna │ 0 │ 0 │ 0 │ 0 │ 0 │ ... 6 more │
├────────────────────┼────────────────────┼────────────────────────────────┼───────────┼───────────┼──────────┼───────────┼───────────┼──────────────┤
│ Ribosomal_S6_30904 │ 92.5 │ (s) Anaerococcus sp002359915 │ 0.528958 │ 0 │ 0.14157 │ 0 │ 0 │ ... 6 more │
╘════════════════════╧════════════════════╧════════════════════════════════╧═══════════╧═══════════╧══════════╧═══════════╧═══════════╧══════════════╛
Almost instantaneously, we learn about the coverages of a given SCG across samples.
You can always use --output-file parameter to store the full output into a TAB-delmited file.
Contigs db + profile db + collection
When we have a metagenomic assembly, in most cases we either perform manual binning and store our bins in a collection, or use an external binning algorithm to import a collection using the program anvi-import-collection into our profile database. In the IDG there is already a collection that describes genome bins we had identified in this dataset, so I will import it into the profile database to demonstrate how collections are used in SCG taxonomy:
anvi-import-collection additional-files/collections/merens.txt \
-p PROFILE.db \
-c CONTIGS.db \
-C COLLECTION
Show/hide Displaying collections in a profile database
And you can always check collections in your profile databases easily. For instance this is what I get after importing the collection COLLECTION:
anvi-show-collections-and-bins -p PROFILE.db
Collection: "CONCOCT"
===============================================
Collection ID ................................: CONCOCT
Number of bins ...............................: 34
Number of splits described ...................: 4,784
Bin names ....................................: Bin_1, Bin_10, Bin_11, Bin_12, Bin_13, Bin_14, Bin_15, Bin_16, Bin_17, Bin_18, Bin_19, Bin_2, Bin_20, Bin_21, Bin_22, Bin_23, Bin_24, Bin_25, Bin_26, Bin_27, Bin_28, Bin_29, Bin_3, Bin_30, Bin_31, Bin_32, Bin_33, Bin_34, Bin_4, Bin_5, Bin_6, Bin_7, Bin_8, Bin_9
Collection: "COLLECTION"
===============================================
Collection ID ................................: COLLECTION
Number of bins ...............................: 13
Number of splits described ...................: 4,451
Bin names ....................................: Aneorococcus_sp, C_albicans, E_facealis, F_magna, L_citreum, P_acnes, P_avidum, P_rhinitidis, S_aureus, S_epidermidis, S_hominis, S_lugdunensis, Streptococcus
The bin names in this collection are taxonomic affiliations for these bins as they appeared in the original publication of this dataset by Sharon et al., so they can serve as ground truths for this little experiment. Now we can run the scg-taxonomy estimation on this collection:
anvi-estimate-scg-taxonomy -c CONTIGS.db \
-p PROFILE.db \
-C COLLECTION
which returns this:
╒═════════════════╤══════════════╤═══════════════════╤══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╕
│ │ total_scgs │ supporting_scgs │ taxonomy │
╞═════════════════╪══════════════╪═══════════════════╪══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╡
│ P_rhinitidis │ 22 │ 20 │ Bacteria / Firmicutes / Clostridia / Tissierellales / Peptoniphilaceae / Peptoniphilus / Peptoniphilus rhinitidis │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ P_avidum │ 22 │ 18 │ Bacteria / Actinobacteriota / Actinobacteria / Propionibacteriales / Propionibacteriaceae / Cutibacterium / Cutibacterium avidum │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ E_facealis │ 21 │ 20 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Enterococcaceae / Enterococcus / Enterococcus faecalis │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ S_epidermidis │ 21 │ 15 │ Bacteria / Firmicutes / Bacilli / Staphylococcales / Staphylococcaceae / Staphylococcus / Staphylococcus epidermidis │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ L_citreum │ 17 │ 17 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Lactobacillaceae / Leuconostoc / Leuconostoc citreum │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ F_magna │ 15 │ 15 │ Bacteria / Firmicutes / Clostridia / Tissierellales / Helcococcaceae / Finegoldia / Finegoldia magna │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ S_aureus │ 15 │ 15 │ Bacteria / Firmicutes / Bacilli / Staphylococcales / Staphylococcaceae / Staphylococcus / │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ S_hominis │ 15 │ 10 │ Bacteria / Firmicutes / Bacilli / Staphylococcales / Staphylococcaceae / Staphylococcus / Staphylococcus hominis │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Streptococcus │ 10 │ 10 │ Bacteria / Firmicutes / Bacilli / Lactobacillales / Streptococcaceae / Streptococcus / │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ Aneorococcus_sp │ 9 │ 6 │ Bacteria / Firmicutes / Clostridia / Tissierellales / Helcococcaceae / Anaerococcus / Anaerococcus sp002359915 │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ S_lugdunensis │ 3 │ 2 │ Bacteria / Firmicutes / Bacilli / Staphylococcales / Staphylococcaceae / Staphylococcus / Staphylococcus lugdunensis │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ C_albicans │ 0 │ 0 │ / / / / / / │
├─────────────────┼──────────────┼───────────────────┼──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────┤
│ P_acnes │ 0 │ 0 │ / / / / / / │
╘═════════════════╧══════════════╧═══════════════════╧══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════╛
Everything seems to be working well (with some exceptions).
It is important to remember two things: One, this is a simple human gut metagenome and for your environmental metagenomes things will not be as smooth (but those of us who study environmental metagenomes are already aware of that). Two, please note the last two lines. Why are they blank? If you were to look at the completion estimates of the bins in this collection,
anvi-estimate-genome-completeness -p PROFILE.db \
-c CONTIGS.db \
-C COLLECTION
This is what I get:
╒═════════════════╤══════════╤══════════════╤════════════════╤════════════════╤══════════════╤════════════════╕
│ bin name │ domain │ confidence │ % completion │ % redundancy │ num_splits │ total length │
╞═════════════════╪══════════╪══════════════╪════════════════╪════════════════╪══════════════╪════════════════╡
│ S_lugdunensis │ BACTERIA │ 0.3 │ 39.44 │ 2.82 │ 227 │ 2302326 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ P_rhinitidis │ BACTERIA │ 0.9 │ 94.37 │ 0 │ 85 │ 1796165 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ Aneorococcus_sp │ BACTERIA │ 0.3 │ 32.39 │ 5.63 │ 495 │ 836796 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ F_magna │ BACTERIA │ 0.6 │ 60.56 │ 2.82 │ 516 │ 1002952 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ P_acnes │ BLANK │ 1 │ 0 │ 0 │ 206 │ 290126 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ L_citreum │ BACTERIA │ 0.8 │ 71.83 │ 2.82 │ 551 │ 1231473 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ E_facealis │ BACTERIA │ 1 │ 100 │ 4.23 │ 140 │ 2865861 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ Streptococcus │ BACTERIA │ 0.4 │ 36.62 │ 1.41 │ 395 │ 659098 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ S_hominis │ BACTERIA │ 0.6 │ 78.87 │ 1.41 │ 135 │ 2135371 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ C_albicans │ EUKARYA │ 0.7 │ 83.13 │ 7.23 │ 1236 │ 13609943 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ S_epidermidis │ BACTERIA │ 0.9 │ 97.18 │ 2.82 │ 194 │ 2572205 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ S_aureus │ BACTERIA │ 0.7 │ 81.69 │ 2.82 │ 140 │ 2748498 │
├─────────────────┼──────────┼──────────────┼────────────────┼────────────────┼──────────────┼────────────────┤
│ P_avidum │ BACTERIA │ 1 │ 98.59 │ 0 │ 131 │ 2506801 │
╘═════════════════╧══════════╧══════════════╧════════════════╧════════════════╧══════════════╧════════════════╛
Which shows that the bin P_acnes has no completion estimate. It doesn’t have a completion estimate, because it doesn’t have a high enough number of SCGs (according to anvi’o) to even make a suggestion regarding how complete it is. In contrast, C_albicans has a fairly good completion score. But even though the BUSCO HMMs Tom Delmont had curated for anvi’o predict its completion well, this bin is a eukaryotic bin. Sadly, scg-taxonomy will not work for eukaryotic genomes (until GTDB starts considering them).
Since we have a profile database, we could also estimate coverages of these bins by simply adding --compute-scg-coverages flag to our command. And expectedly, when I run this,
anvi-estimate-scg-taxonomy -c CONTIGS.db \
-p PROFILE.db \
-C COLLECTION \
--compute-scg-coverages
This is what I get:
╒═════════════════╤══════════════╤═══════════════════╤════════════════════════════════╤═══════════╤═══════════╤════════════╤═══════════╤═══════════╤══════════════╕
│ │ total_scgs │ supporting_scgs │ taxonomy │ DAY_15A │ DAY_15B │ DAY_16 │ DAY_17A │ DAY_17B │ ... 6 more │
╞═════════════════╪══════════════╪═══════════════════╪════════════════════════════════╪═══════════╪═══════════╪════════════╪═══════════╪═══════════╪══════════════╡
│ P_rhinitidis │ 22 │ 20 │ (s) Peptoniphilus rhinitidis │ 0 │ 0.0889543 │ 0.00294413 │ 0 │ 0.089591 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ P_avidum │ 22 │ 18 │ (s) Cutibacterium avidum │ 12.5869 │ 3.95421 │ 2.42292 │ 0 │ 0 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ E_facealis │ 21 │ 20 │ (s) Enterococcus faecalis │ 309.381 │ 602.743 │ 604.223 │ 153.103 │ 963.167 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ S_epidermidis │ 21 │ 15 │ (s) Staphylococcus epidermidis │ 107.146 │ 175.284 │ 143.184 │ 27.0741 │ 119.61 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ L_citreum │ 17 │ 17 │ (s) Leuconostoc citreum │ 0.0664338 │ 0.02886 │ 1.90741 │ 0 │ 0.374945 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ S_aureus │ 15 │ 15 │ (g) Staphylococcus │ 0.131936 │ 0.871665 │ 0.962612 │ 0.0687208 │ 0.553133 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ F_magna │ 15 │ 15 │ (s) Finegoldia magna │ 0 │ 0 │ 0 │ 0 │ 0 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ S_hominis │ 15 │ 10 │ (s) Staphylococcus hominis │ 2.75788 │ 15.9922 │ 10.1896 │ 0.0100202 │ 9.60573 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ Streptococcus │ 10 │ 10 │ (g) Streptococcus │ 0 │ 0 │ 0 │ 0 │ 0 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ Aneorococcus_sp │ 9 │ 6 │ (s) Anaerococcus sp002359915 │ 1.18094 │ 0.61353 │ 0.526029 │ 0 │ 0 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ S_lugdunensis │ 3 │ 2 │ (s) Staphylococcus lugdunensis │ 0 │ 4.61847 │ 5.10752 │ 0 │ 0 │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ P_acnes │ 0 │ 0 │ (NA) NA │ │ │ │ │ │ ... 6 more │
├─────────────────┼──────────────┼───────────────────┼────────────────────────────────┼───────────┼───────────┼────────────┼───────────┼───────────┼──────────────┤
│ C_albicans │ 0 │ 0 │ (NA) NA │ │ │ │ │ │ ... 6 more │
╘═════════════════╧══════════════╧═══════════════════╧════════════════════════════════╧═══════════╧═══════════╧════════════╧═══════════╧═══════════╧══════════════╛
Many contigs dbs for many metagenomes
So far we have worked on individual contigs databases that describe a single genome or a single metagenome (with or without associated profile databases). But what if if you have many contigs databases? The program anvi-estimate-scg-taxonomy can work with multiple contigs databases, but to tell anvi’o about where to find those files, we will use a special file format.
As you may already know, anvi’o pangenomics workflow uses two file formats to describe internal and external genomes. Just a reminder, external genomes file looks like this:
| name | contigs_db_path |
|---|---|
| Genome_Name_01 | /path/to/contigs-01.db |
| Genome_Name_02 | /path/to/contigs-02.db |
| Genome_Name_03 | /path/to/contigs-03.db |
| (…) | (…) |
Where each contig database is listed with a name. The expectation for the external genomes file is that each contigs database represents a single genome. The internal genomes file, in contrast, describes genomes defined in anvi’o collections:
| name | bin_id | collection_id | profile_db_path | contigs_db_path |
|---|---|---|---|---|
| Genome_Name_01 | Bin_id_01 | Collection_A | /path/to/profile.db | /path/to/contigs.db |
| Genome_Name_02 | Bin_id_02 | Collection_A | /path/to/profile.db | /path/to/contigs.db |
| Genome_Name_03 | Bin_id_03 | Collection_B | /path/to/another_profile.db | /path/to/another/contigs.db |
| (…) | (…) | (…) | (…) | (…) |
Again, each entry in internal genomes files is also expect to be a single genome. You can indeed try to trick anvi’o, but you shouldn’t. Even the most carefully implemented and tested software rarely work, but software you trick will almost never do.
The format of the metagenomes file is similar to the format of external genomes file, but there are two differences. One, each contigs database in each entry in a metagenomes file would be considered to be a metagenome, rather than a genome. And two, for each contigs database the user can optionally list a single profile database (i.e., the output of the anvi-profile and not anvi-merge):
| name | contigs_db_path | profile_db_path |
|---|---|---|
| Metagenome_Name_01 | /path/to/contigs-01.db | /path/to/profile-01.db |
| Metagenome_Name_02 | /path/to/contigs-02.db | /path/to/profile-02.db |
| Metagenome_Name_03 | /path/to/contigs-03.db | /path/to/profile-03.db |
| (…) | (…) | (…) |
So without the optional column profile_db_path, the metagenomes file is identical to the external genomes file. But as I mentioned, external genomes file is meant to describe genomes, and metagenomes file is meant to describe metagenomes.
Let’s say we have a bunch of metagenomes from the Human Microbiome Project, and our metagenomes file looks like this:
| name | contigs_db_path | profile_db_path |
|---|---|---|
| USA0001 | USA0001_01-contigs.db | USA0001_01/USA0001_01/PROFILE.db |
| USA0003 | USA0003_01-contigs.db | USA0003_01/USA0003_01/PROFILE.db |
| USA0004 | USA0004_01-contigs.db | USA0004_01/USA0004_01/PROFILE.db |
| USA0005 | USA0005_01-contigs.db | USA0005_01/USA0005_01/PROFILE.db |
| USA0006 | USA0006_01-contigs.db | USA0006_01/USA0006_01/PROFILE.db |
| USA0007 | USA0007_01-contigs.db | USA0007_01/USA0007_01/PROFILE.db |
| USA0008 | USA0008_01-contigs.db | USA0008_01/USA0008_01/PROFILE.db |
| USA0009 | USA0009_01-contigs.db | USA0009_01/USA0009_01/PROFILE.db |
Here, for each line, name is a unique name for the entry, contigs_db_path is the anvi’o contigs database generated from the assembly of name, and profile_db_path is what we got back from the program anvi-profile when we recruited reads from the metagenome using our assembly. When I run the following command,
anvi-estimate-scg-taxonomy --metagenomes metagenomes.txt \
--output-file-prefix METAGENOMES-EXAMPLE
Anvi’o goes through all contigs databases, finds the frequencies of each SCG, and picks one that is the most frequent across them. This is what my terminal says:
(...)
Long-format output ...........................: METAGENOMES-EXAMPLE-LONG-FORMAT.txt
The output file will conveniently contain all taxonomic levels, and their coverages in each metagenome. For brevity, here let’s take a look at the phylum level content. Running this,
grep t_phylum METAGENOMES-EXAMPLE-LONG-FORMAT.txt | sort -k 3
Gives me this:
| entry_id | metagenome_name | taxon | coverage | taxonomic_level |
|---|---|---|---|---|
| (…) | (…) | (…) | (…) | (…) |
| 22 | USA0003 | Bacteroidota | 1034.4713634221948 | t_phylum |
| 41 | USA0007 | Bacteroidota | 1615.1358016297484 | t_phylum |
| 31 | USA0005 | Bacteroidota | 2255.2101597095934 | t_phylum |
| 26 | USA0004 | Bacteroidota | 2297.481202094419 | t_phylum |
| 51 | USA0009 | Bacteroidota | 2565.7820609024084 | t_phylum |
| 37 | USA0006 | Bacteroidota | 443.17724125228267 | t_phylum |
| 46 | USA0008 | Bacteroidota | 484.36157980489156 | t_phylum |
| 16 | USA0001 | Bacteroidota | 932.4783076099221 | t_phylum |
| 47 | USA0008 | Firmicutes | 0.0 | t_phylum |
| 17 | USA0001 | Firmicutes | 126.83701119138941 | t_phylum |
| 27 | USA0004 | Firmicutes | 1470.5449459499075 | t_phylum |
| 21 | USA0003 | Firmicutes | 1602.1891603200138 | t_phylum |
| 42 | USA0007 | Firmicutes | 168.53206707580267 | t_phylum |
| 36 | USA0006 | Firmicutes | 2272.792244006277 | t_phylum |
| 52 | USA0009 | Firmicutes | 429.89995475707457 | t_phylum |
| 32 | USA0005 | Firmicutes | 704.7388912444274 | t_phylum |
| 39 | USA0006 | Proteobacteria | 0.0 | t_phylum |
| 48 | USA0008 | Proteobacteria | 0.0 | t_phylum |
| 53 | USA0009 | Proteobacteria | 0.0 | t_phylum |
| 29 | USA0004 | Proteobacteria | 11.738617200674536 | t_phylum |
| 18 | USA0001 | Proteobacteria | 17.961693548387096 | t_phylum |
| 33 | USA0005 | Proteobacteria | 30.329506731467518 | t_phylum |
| 43 | USA0007 | Proteobacteria | 7.588628762541806 | t_phylum |
| 24 | USA0003 | Proteobacteria | 9.264563106796116 | t_phylum |
| 19 | USA0001 | Unknown_phyla | 0.0 | t_phylum |
| 30 | USA0004 | Unknown_phyla | 0.0 | t_phylum |
| 40 | USA0006 | Unknown_phyla | 0.0 | t_phylum |
| 44 | USA0007 | Unknown_phyla | 0.0 | t_phylum |
| 49 | USA0008 | Unknown_phyla | 0.0 | t_phylum |
| 54 | USA0009 | Unknown_phyla | 0.0 | t_phylum |
| 35 | USA0005 | Unknown_phyla | 3.648562300319489 | t_phylum |
| 25 | USA0003 | Unknown_phyla | 9.177676537585421 | t_phylum |
| 20 | USA0001 | Verrucomicrobiota | 0.0 | t_phylum |
| 45 | USA0007 | Verrucomicrobiota | 0.0 | t_phylum |
| 50 | USA0008 | Verrucomicrobiota | 0.0 | t_phylum |
| 55 | USA0009 | Verrucomicrobiota | 0.0 | t_phylum |
| 34 | USA0005 | Verrucomicrobiota | 21.08811188811189 | t_phylum |
| 23 | USA0003 | Verrucomicrobiota | 22.668446982908964 | t_phylum |
| 28 | USA0004 | Verrucomicrobiota | 42.04647160068847 | t_phylum |
| 38 | USA0006 | Verrucomicrobiota | 9.919685039370078 | t_phylum |
| (…) | (…) | (…) | (…) | (…) |
If this output is not what you want, you can also try the --matrix-format flag:
anvi-estimate-scg-taxonomy --metagenomes metagenomes.txt \
--output-file-prefix METAGENOMES-EXAMPLE \
--matrix-format
This time, the output files are reported this way:
(...)
Output matrix for "t_domain" .................: METAGENOMES-EXAMPLE-t_domain-MATRIX.txt
Output matrix for "t_phylum" .................: METAGENOMES-EXAMPLE-t_phylum-MATRIX.txt
Output matrix for "t_class" ..................: METAGENOMES-EXAMPLE-t_class-MATRIX.txt
Output matrix for "t_order" ..................: METAGENOMES-EXAMPLE-t_order-MATRIX.txt
Output matrix for "t_family" .................: METAGENOMES-EXAMPLE-t_family-MATRIX.txt
Output matrix for "t_genus" ..................: METAGENOMES-EXAMPLE-t_genus-MATRIX.txt
Output matrix for "t_species" ................: METAGENOMES-EXAMPLE-t_species-MATRIX.txt
As you can imagine, the content of the matrix file for phylum looks like this:
| taxon | USA0001 | USA0003 | USA0004 | USA0005 | USA0006 | USA0007 | USA0008 | USA0009 |
|---|---|---|---|---|---|---|---|---|
| Bacteroidota | 932.48 | 1034.47 | 2297.48 | 2255.21 | 443.18 | 1615.14 | 484.36 | 2565.78 |
| Firmicutes | 126.84 | 1602.19 | 1470.54 | 704.74 | 2272.79 | 168.53 | 0.00 | 429.90 |
| Proteobacteria | 17.96 | 9.26 | 11.74 | 30.33 | 0.00 | 7.59 | 0.00 | 0.00 |
| Unknown_phyla | 0.00 | 9.18 | 0.00 | 3.65 | 0.00 | 0.00 | 0.00 | 0.00 |
| Verrucomicrobiota | 0.00 | 22.67 | 42.05 | 21.09 | 9.92 | 0.00 | 0.00 | 0.00 |
Tadaa! 🎉🥳
All reported values are raw coverages for genes that are used. But if you want anvi’o also to report percent normalized values for convenience, drop us a line (we are too lazy to do it, but we are too nice to say no to people).
Estimating taxonomy in the interface
In other words, the fun stuff.
We embarked upon this journey because we needed something extremely fast and relatively accurate while working with metagenomes. Identifying a genome bin in anvi’o is extremely easy, but figuring out the taxonomy of these bins in the interface has not been possible. The scg-taxonomy subsystem bridges that gap now. For instance, if I initiate the interactive interface for the infant gut metagenome,
anvi-interactive -p PROFILE.db \
-c CONTIGS.db
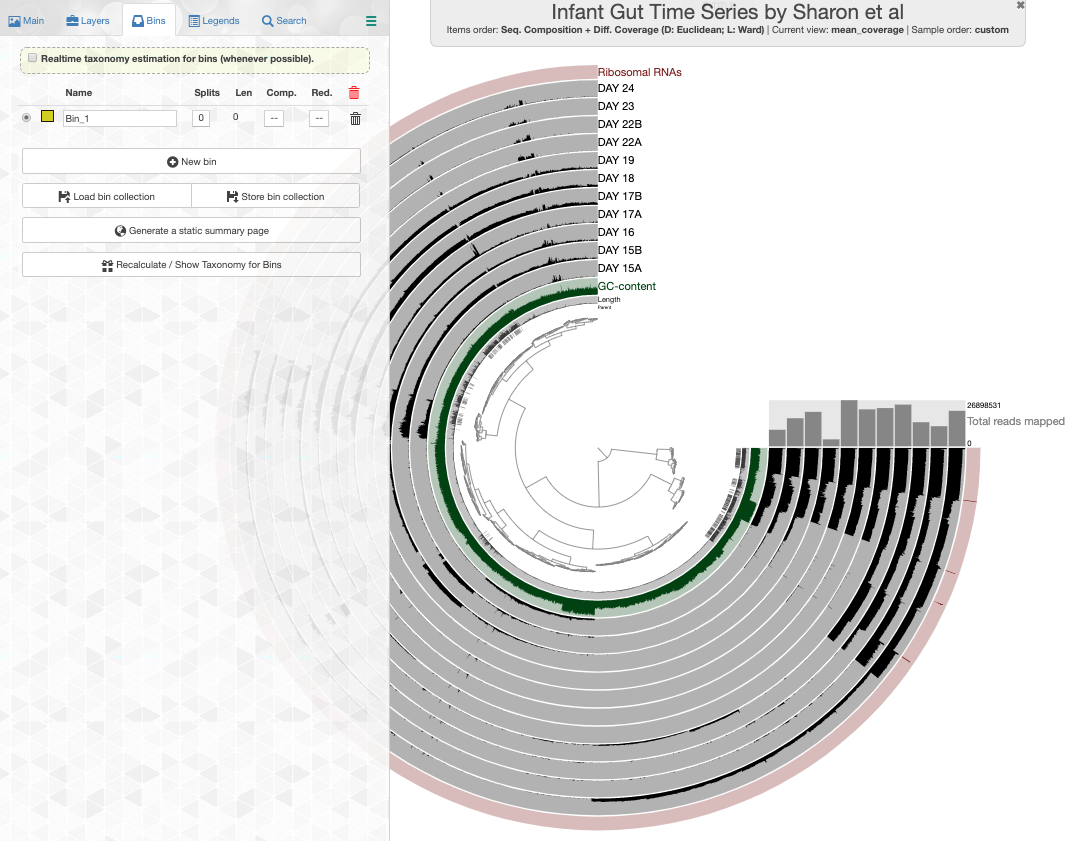
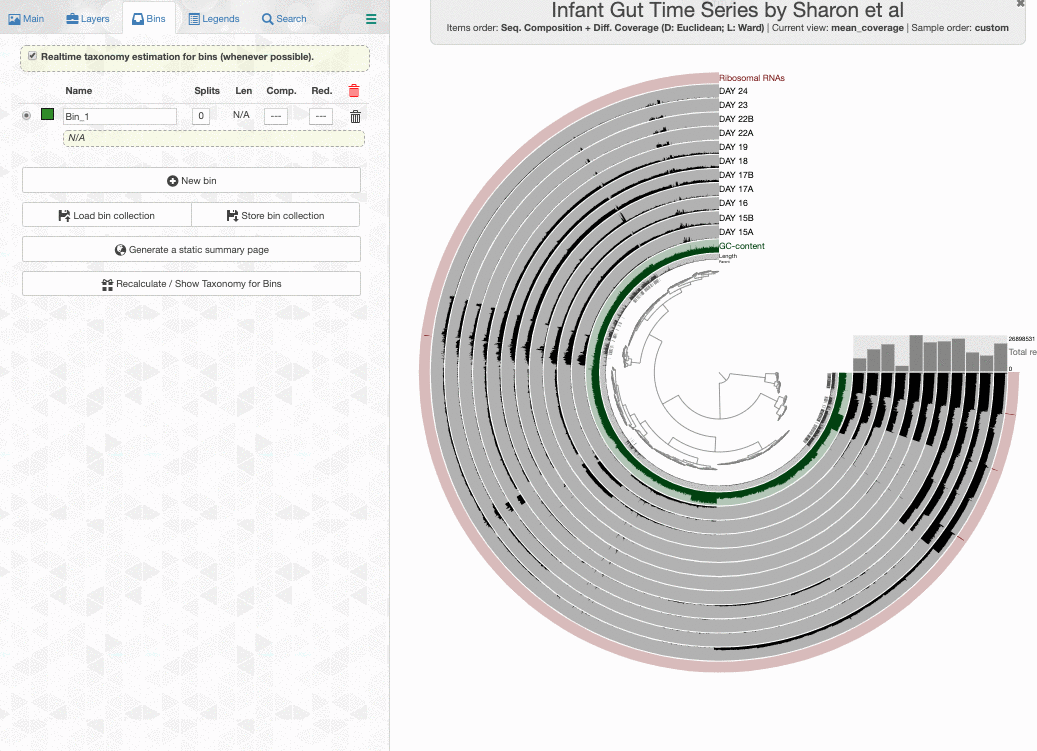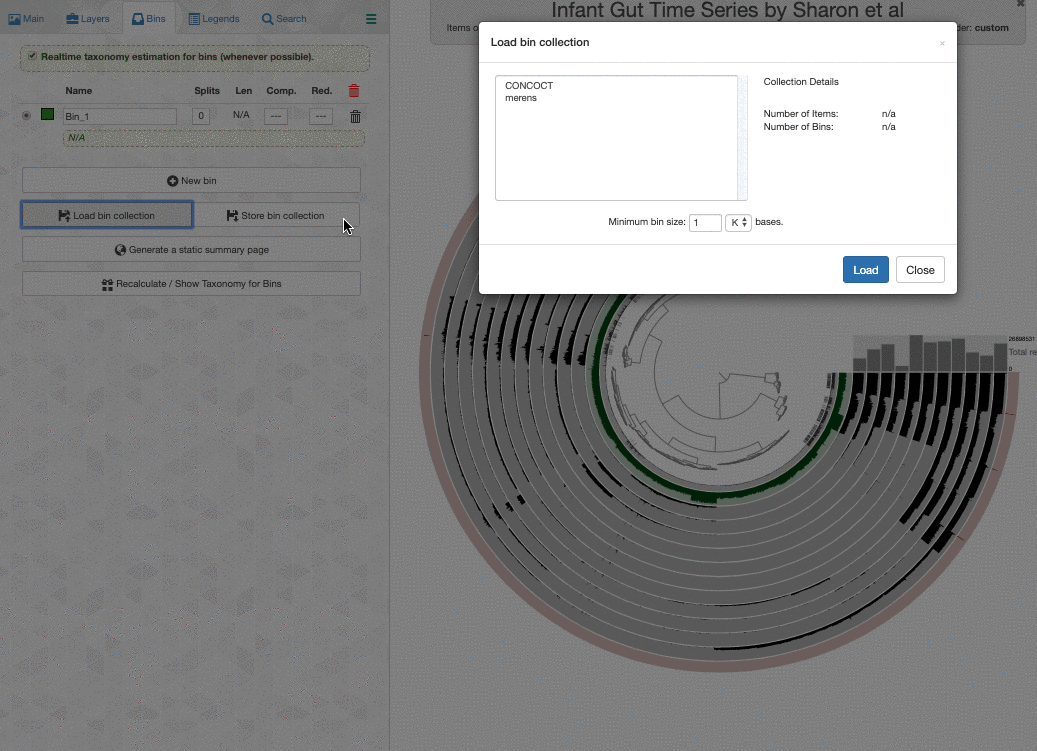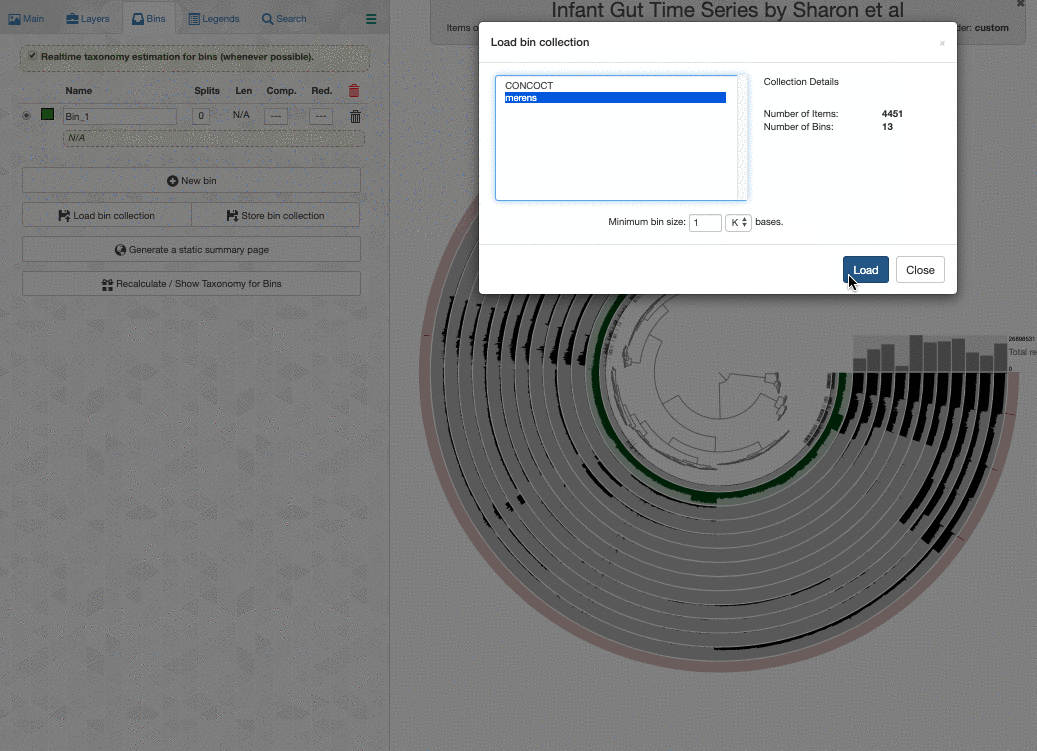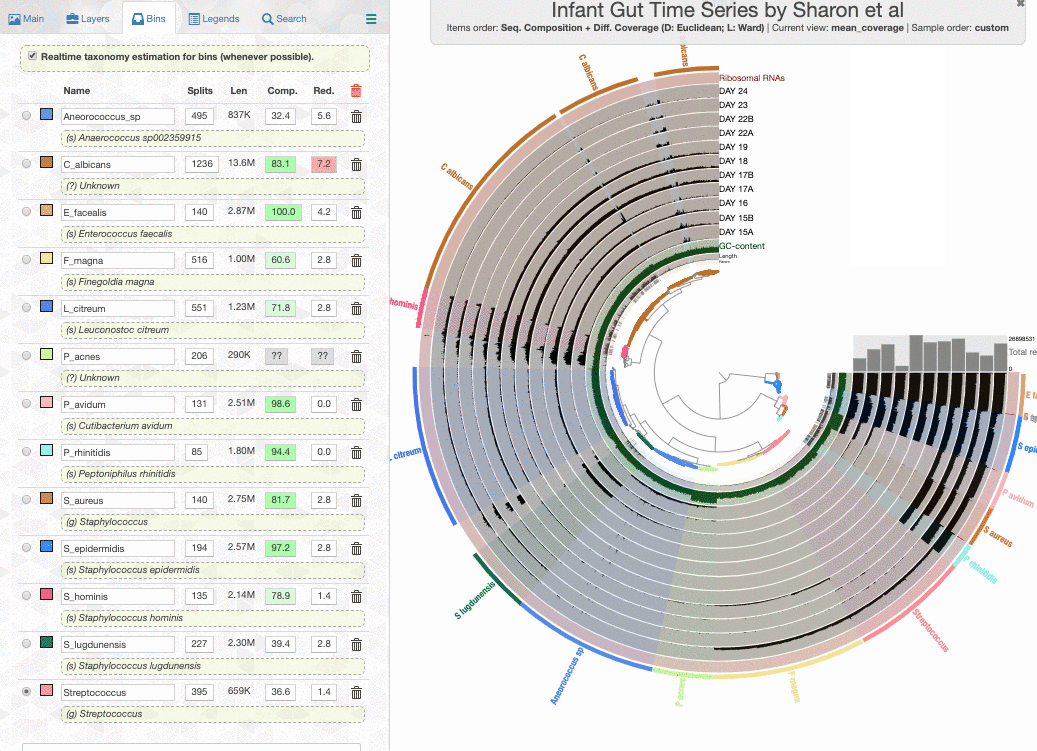
click the draw button, and switch to the Bin panel, I will see this display:
Look what happens if I enable the real-time taxonomy estimation for my bins:
Well. I don’t know how to put this, but this is real-time. This is how fast a new set of contigs are affiliated with taxonomy in the interface.
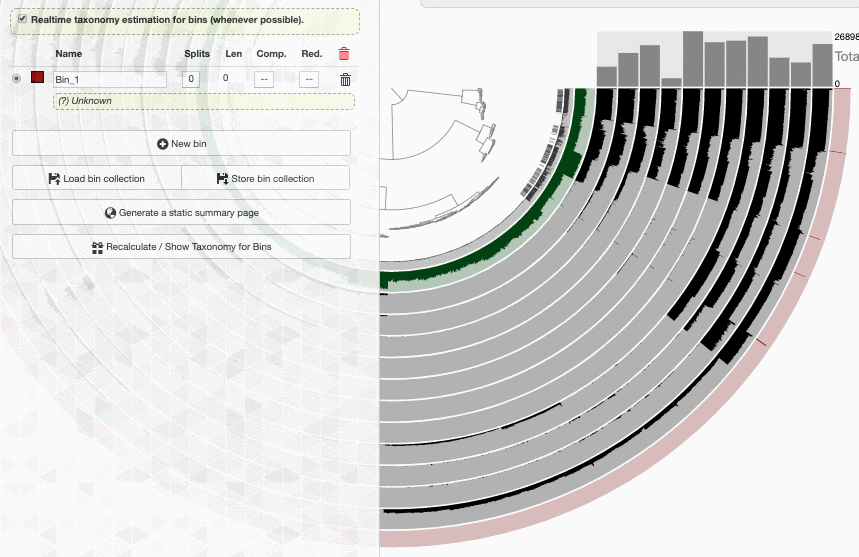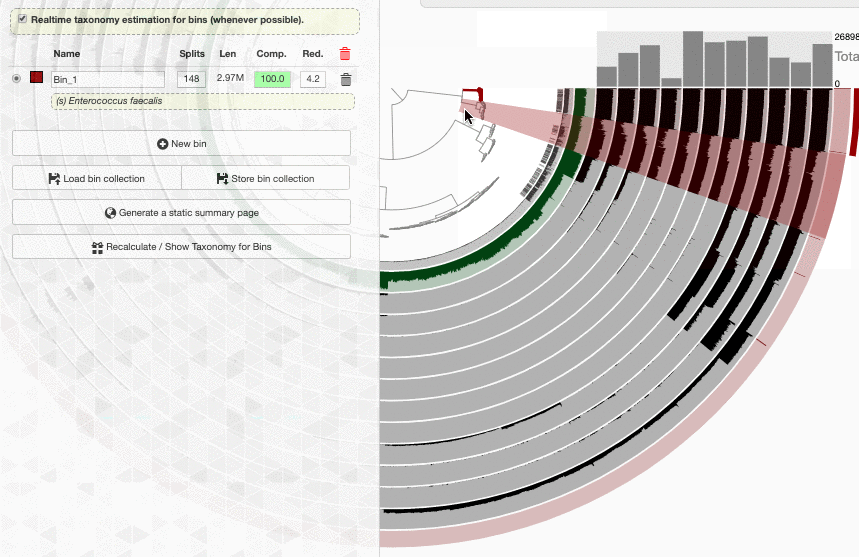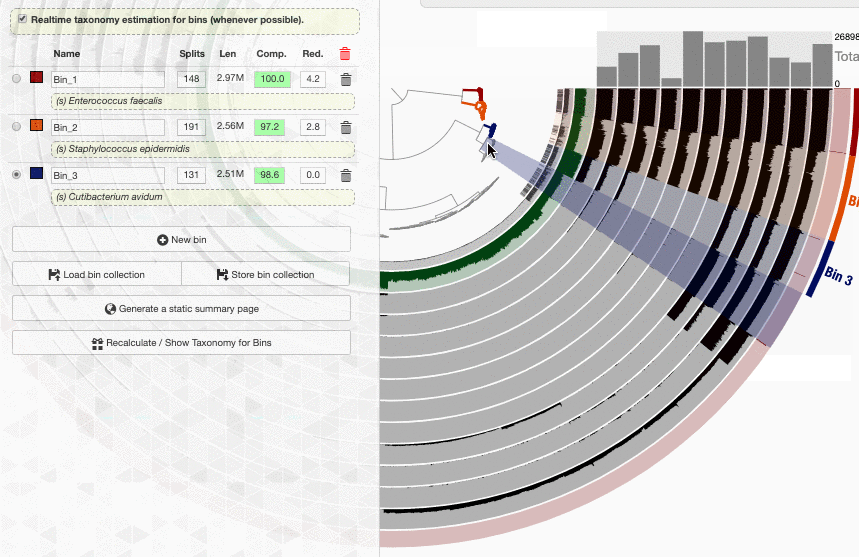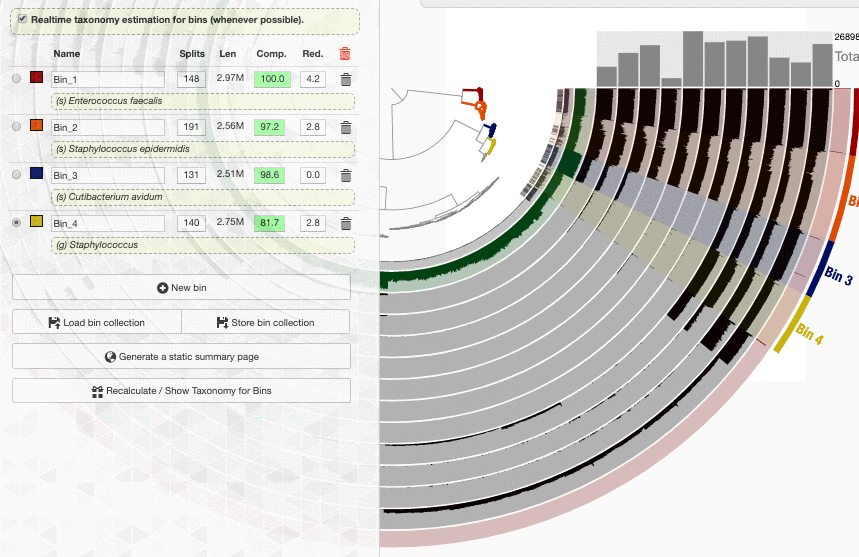
Here is what happens when I load the colleciton merens to this interface:
It is happening so rapidly, it looks as if anvi’o knew the taxonomy for these bins. But in reality the taxonomy shown underneath bin names are computed on the fly.
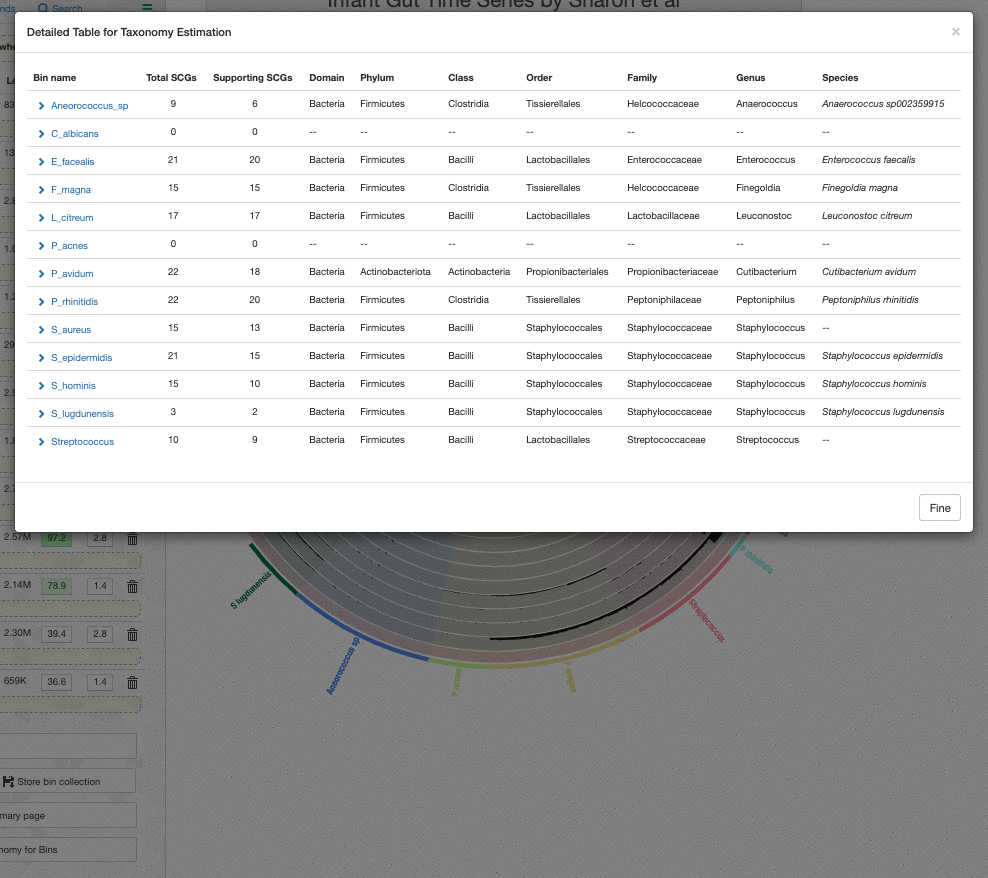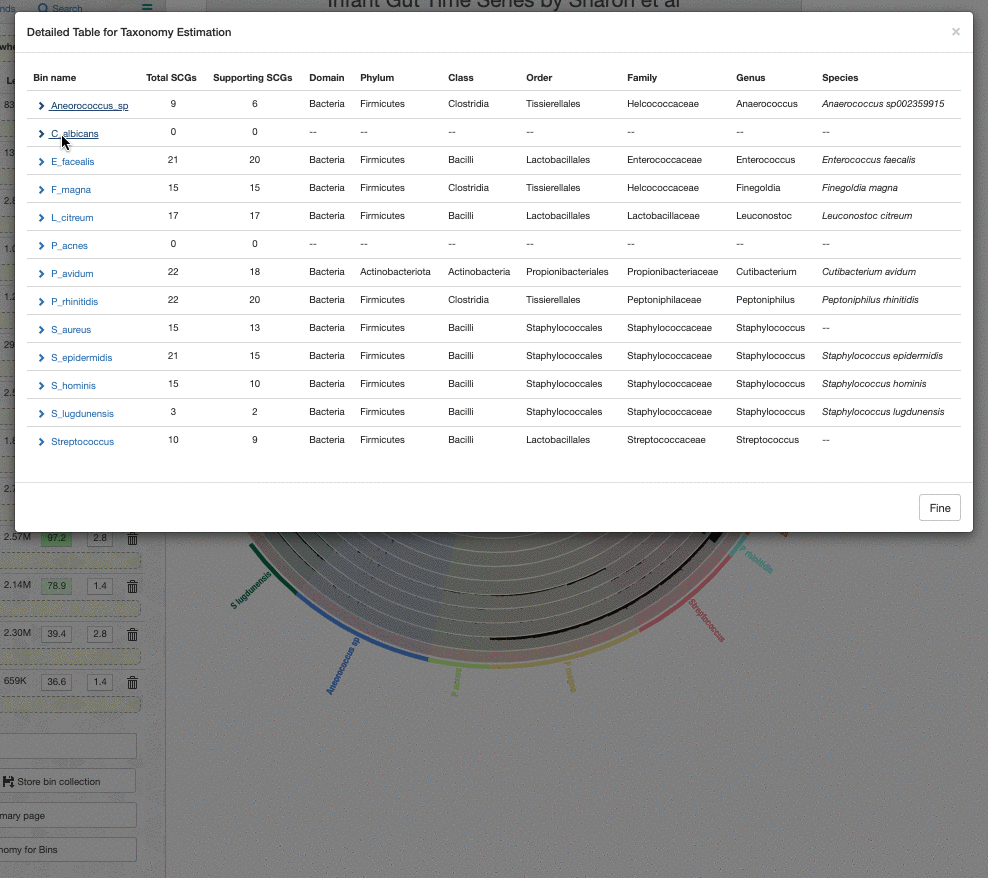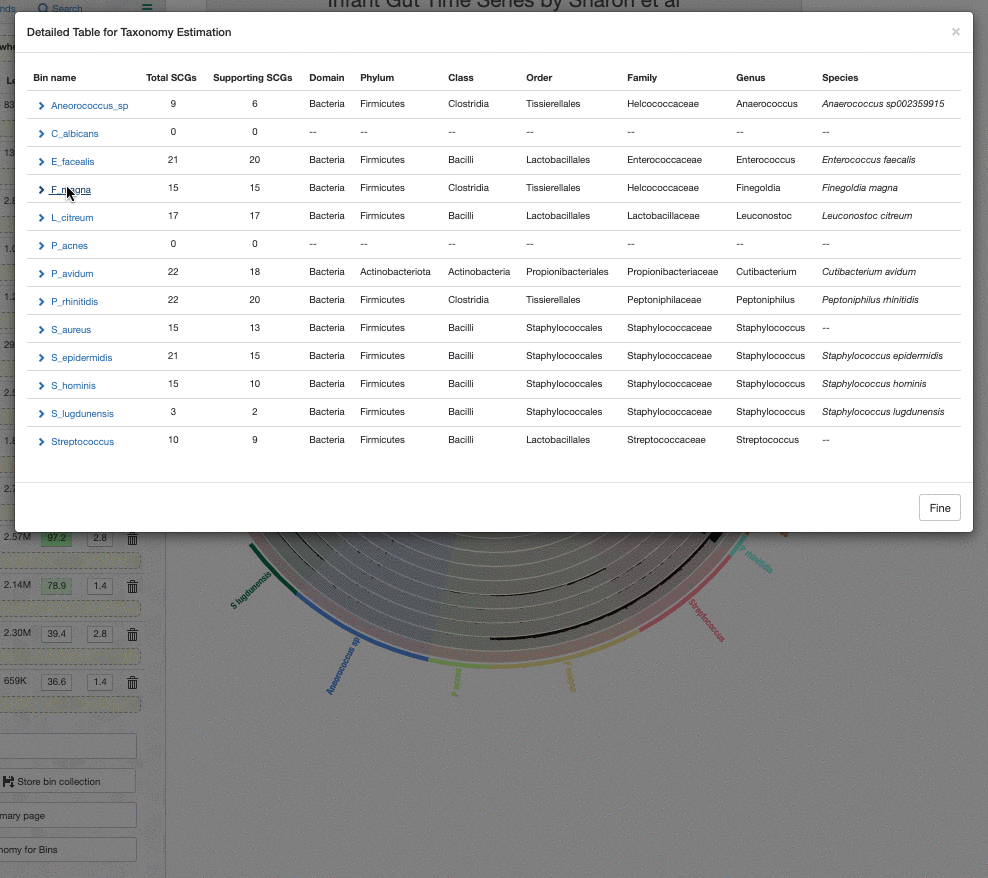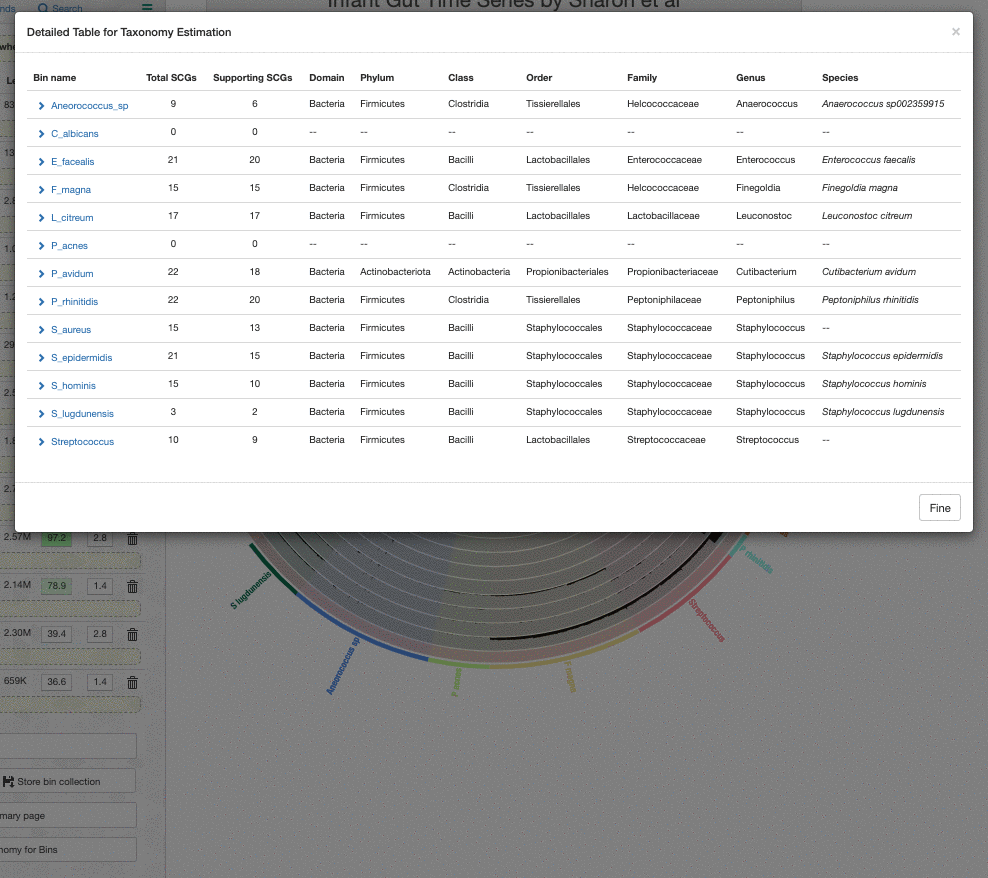
One thing to remember is that the taxonomy information shown on the bins menu is a consensus taxonomy calculated from the taxonomic affiliations of all relevant SCGs in a given bin. It is possible to see the underlying data for this through the taxonomy table interface, which you can open by clicking on the “Recalculate / Show Taxonomy for Bins” button in the Bins panel. When you click that, you see the following table that shows the consensus taxonomy, and the SCGs from which it was computed:
Final words and some warnings
You will not get perfect taxonomy with scg-taxonomy, but you will some taxonomy very rapidly. Anvi’o scg-taxonomy module will not estimate taxonomy for bins that are very low completion. It will not estimate taxonomy for eukaryotic bins. In --metagenome-mode you will not see any evidence of the populations from which the assembler did not recover any of the ribosomal proteins anvi’o uses for taxonomy estimation.
Yes. You will not get perfect taxonomy, but the reality is nothing will give you perfect taxonomy. There is no such thing as perfect taxonomy. Taxonomy is a mess. Thanks to heroic efforts by those who try to find an order in this mess (such as those who gave us SILVA or GTDB), we, as a community, try to find the best place to attach our genomes in the tree of life. But from building trees to determining single-copy core genes, to identifying them in genomes, to defining and assigning taxonomic names, everything slowly falls apart as we struggle between sensitivity, feasibility, and accuracy of our approaches. For those of us who need taxonomy, the struggle is real. Our best option is to make sure we confirm the taxonomic affiliations of our genomes and bins. Which, despite the popular belief, may require manual work in some occasions using pangenomics, phylogenomics, studing ANI, BLAST’ing things around, etc. But I still hope the anvi’o scg-taxonomy will help you to develop some initial insights into your taxonomic mess 😇
I would also suggest you to take look at GTDB-Tk. It will almost certainly do a better job to estimate taxonomy (we are hoping to publish a small note with proper benchmarks on accuracy). The GitHub page of GTDB-Tk suggests that it requires ~100Gb of memory, ~27Gb of storage, and takes about ~1 hour per 1,000 genomes when using 64 CPUs. Anvi’o scg-taxonomy takes about 100Mb space on a laptop computer with 16Gb memory and 4 CPUs, and takes about 20 seconds to assign taxonomy to 1,000 genomes. Of course this time doesn’t include standard items in anvi’o workflow that take place before anvi’o scg-taxonomy but are required for it to run, such as the time it takes to create an anvi’o contigs database, running HMMs on it, etc. But regardless, you should always be extra careful with things that are fast.
Both for anvi-run-scg-taxonomy and anvi-estimate-scg-taxonomy programs accept --debug flags. If things don’t make sense, you should re-run these programs with the --debug flag to see additional information.
This is an extremely new feature and we hope it will contribute to your journey in microbial ‘omics and your anvi’o experience. Please keep in mind just like everything in anvi’o, the evolution of this feature will be heavily influenced by your feedback and suggestions. Please let us know if something doesn’t work for you!
If you find a mistake on this page or would you like to update something in it, please feel free to edit its source by clicking the edit button at the top-right corner (which you will see if you are logged in to GitHub) 😇